 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Manes.09G001400.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Manihoteae; Manihot
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 844aa MW: 96486.8 Da PI: 7.1992 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 84.6 | 1.6e-26 | 85 | 187 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk............tekerrtraetrtgCkaklkvkkekdgkwev 78
+fY+eYA+++GF++ +++s++sk+++e+++++f Cs++g+++e +k+ e+ +r+ ++t+Cka+++vk++ dgkw++
Manes.09G001400.2.p 85 SFYQEYARSTGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYDKSfnrprarqnkqdPENGTGRRSCSKTDCKASMHVKRRPDGKWVI 174
5*******************************************999999998776555444555999********************** PP
FAR1 79 tkleleHnHelap 91
+++++eHnHel p
Manes.09G001400.2.p 175 HSFVKEHNHELLP 187
**********975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.9E-24 | 85 | 187 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 7.7E-26 | 285 | 376 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.322 | 565 | 601 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 3.2E-6 | 576 | 603 | IPR006564 | Zinc finger, PMZ-type |
| Pfam | PF04434 | 1.4E-4 | 577 | 600 | IPR007527 | Zinc finger, SWIM-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 844 aa Download sequence Send to blast |
MDIDLRLPSG EHDKDNEEPN GIDNMLSEDK LHNGDVGTEN IVNVAEEVHA IEGGPMCSPT 60 TEFKEEANLE PLAGMEFESH GAAYSFYQEY ARSTGFNTAI QNSRRSKTSR EFIDAKFACS 120 RYGTKREYDK SFNRPRARQN KQDPENGTGR RSCSKTDCKA SMHVKRRPDG KWVIHSFVKE 180 HNHELLPAQA VSEQTRKMYA AMARQFAEYK NVIGLKNDLK NPFDKGRNSA LEAADAKILL 240 DFFTQLQSLN SNFFYAVELG EDQLLKNLVW IDAKSRYDYV NFCDVVSFDA TYIRSKYKIP 300 LGLFVGVNQH CQFMLLGCAL LSEESTTSYS WLMQTWLRGM GGLAPKVIIT DQDKALKSVI 360 TEVFPNAHHY FFLWSILGKV AENLSQVTKR HENFMAKFEK CIFRSWTNDE FVKRWLKILD 420 RFELRENDVM QSLYEDRDLW VPIYMRDATL AGISTIQRAE SLNSYFDKYL HKKTTVQEFV 480 KQYEAILQDR YEEEAKADSD TWNKQPTLKS PSPLEKSVSG IYTHAVFKKF QVEVLGVVAC 540 HPKMESQDET TVSFRVQDLE KHQDFTVVWN QIRSEVACIC RLYEYKGYLC RHALVVLQMC 600 QQSAIPSQYI LKRWTKDVKS RHLLGEDCEQ VQSRVQRYNL LCQRALKLSE EGSLSQESYN 660 IAFRALEEAF GNCISANNSN KTLAEAGSSA THGLLCIEED NQSRSMNKTN KKKNPAKKRK 720 MNSEQEVTTV GAEDSLQQMD KLRSRAVTLD GYYGAQPSVP GMVQLNLMAP RDNFYGNQQT 780 IQGLGQLNSI APSHDGYYNA QQSMHGLAQM DFFRTPAGFS YGIRQDDPNM RTAQLHDNAS 840 RHA* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
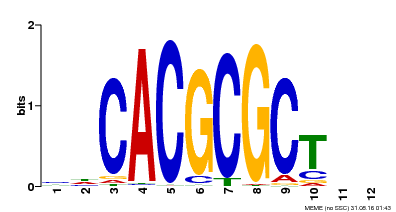
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Manes.09G001400.2.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021622856.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X3 | ||||
| Refseq | XP_021622857.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X3 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A251K2D5 | 0.0 | A0A251K2D5_MANES; Uncharacterized protein | ||||
| STRING | POPTR_0006s02150.1 | 0.0 | (Populus trichocarpa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Manes.09G001400.2.p |



