 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Manes.05G010400.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Manihoteae; Manihot
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 979aa MW: 108393 Da PI: 6.7461 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 131.6 | 2.6e-41 | 158 | 235 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C adls+ak+yhrrhkvC+ hska+++lv +++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Manes.05G010400.1.p 158 VCQVEDCGADLSNAKDYHRRHKVCDLHSKASKALVGNVMQRFCQQCSRFHVLQEFDEGKRSCRRRLAGHNKRRRKTNP 235
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 4.9E-33 | 152 | 220 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.146 | 156 | 233 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.35E-38 | 157 | 237 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 2.4E-29 | 159 | 232 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 4.6E-6 | 755 | 864 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 4.51E-7 | 755 | 860 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 5.14E-7 | 756 | 863 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 979 aa Download sequence Send to blast |
MEARFGGEGE AQAHHFCGMS ATEMRAVGKR SLEWDLNDWK WDGDLFIASP LNPVPSGGMG 60 RQFFPVATGI PVNGNSSNSS SSCSDEVNLG IEQGKRELEK RRRVIVIEDD NLNGEEVGSL 120 SLKLGGHGYP ITQREIRSWE GNSGKKTKLA GGSMSRAVCQ VEDCGADLSN AKDYHRRHKV 180 CDLHSKASKA LVGNVMQRFC QQCSRFHVLQ EFDEGKRSCR RRLAGHNKRR RKTNPEAAGN 240 GSSLNDDQTT SYLLISLLRI LSNMHSNRSD QVTDQDLLSH LLKSLGSHTI EHGGRNISGL 300 LQESRDLLND GNSEQVGHAH GANGANMQTS PVKPSILNNY PAYSEVRDTT VGQVKMNNFD 360 LNDIYVDSDD GAEDIERSPV VTNMMTSSLD CPSWIQQDSH QSSPPQTSRN SDSASAQSPS 420 SSSGDAQSRT DRIIFKLFGK EPNDFPLALR AQILDWLSHS PTDIESYIRP GCVILTIYLR 480 QADATWEELC CNLSSSLGRL LDVSDDAFWR TGWVYIKVQH QVAFVCNGKV VVDTSLSLKS 540 NNCSQILSVK PIAISASEKA QFVIKGINLS RPTTRLLCAV EGKYMFQKDT EELMGRVDNL 600 KGHKELQCVN FSCSIPTVSG RGFIEIEDHG FSSSYFPFIV AEDDVCSDIR MLEGVLEFAE 660 TDADGSGKMV AKNQAMEFIH EIGWLLHRSQ LKSRLDHLDP YTDLFPLKRF KWLMEFSMDH 720 EWCAVVKKLL NILLKGVVGT GEHSSLDLAL SEMGLLHRAV RKNSRSLVEL LLRYAPEKTV 780 PENKLLIGGS HESSLFRPDA TGPAGLTPLH IAAGKDGSED VLDVLTDDPG MVGIEAWKSA 840 RDSTGFTPED YARLRGHYLY IHLVQKKINK RSAVGHVVLD IPEALSDCST KQKQNEEVST 900 SFEIAQTAKR PIQRPCKLCH QKLDYGTAGR SLLYRPAMLS MVAIAAVCVC VALLFKSSPE 960 VVYVFRPFRW ELLDYGTS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-30 | 150 | 232 | 2 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 99 | 103 | KRRRV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
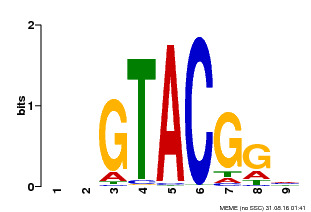
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Manes.05G010400.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021614003.1 | 0.0 | squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A2C9VUD9 | 0.0 | A0A2C9VUD9_MANES; Uncharacterized protein | ||||
| STRING | cassava4.1_000995m | 0.0 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1539 | 34 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Manes.05G010400.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



