 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Manes.01G263300.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Manihoteae; Manihot
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 367aa MW: 38916.3 Da PI: 9.4933 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 120.9 | 1.5e-37 | 84 | 185 | 2 | 104 |
TCP 2 agkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgtssssaseceaesssssasnsssgkaa 91
++k+ +++++hTkv+gR+RR+R++a+caar+F+L++eLG+++d++ti WLl+ a+pai+++tgt++ +a ++ +++ + + s+++++
Manes.01G263300.1.p 84 PPKRASTKDRHTKVEGRGRRIRMPATCAARIFQLTRELGHKSDGETIRWLLEHAEPAIIAATGTGTVPAIAM-SVNGTLKIPTTSNANSE 172
789*************************************************************99999555.22222222222222233 PP
TCP 92 ksaakskksqksa 104
+++++ kk++k++
Manes.01G263300.1.p 173 PNDPSVKKKRKRP 185
3333222222222 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 1.9E-30 | 89 | 172 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 27.141 | 90 | 144 | IPR017887 | Transcription factor TCP subgroup |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008361 | Biological Process | regulation of cell size | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 367 aa Download sequence Send to blast |
MATLIHKQEI EHDDDTRNVD LRISADGDAH TDKLDPTANS IFTTKEVAPK EEPDSEERSP 60 APLGVMPIAV HVPTAIRMPL AAAPPKRAST KDRHTKVEGR GRRIRMPATC AARIFQLTRE 120 LGHKSDGETI RWLLEHAEPA IIAATGTGTV PAIAMSVNGT LKIPTTSNAN SEPNDPSVKK 180 KRKRPANSEY IDISDAAVSV SAPLAPLMTP QPPPPQQQTA TAVIPQGLVP MWAIPSNAVV 240 PGAFFMVPPM AASIAGTPNQ PQIFTFPAAA TPLINISARP ISSFVSSMQQ AANIAVAMPV 300 SSSTISGSKP TKATSVMAPS SSSAPISSST ANSTNTSTTT TQMLRDFSLE IYDKQELQFM 360 TRPSKH* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 177 | 184 | VKKKRKRP |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
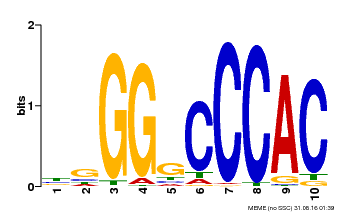
| Motif ID | Method | Source | Motif file |
| MP00667 | PBM | Transfer from GSVIVG01027588001 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Manes.01G263300.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021613812.1 | 0.0 | transcription factor TCP9-like | ||||
| Swissprot | O64647 | 2e-98 | TCP9_ARATH; Transcription factor TCP9 | ||||
| TrEMBL | A0A2C9WRV6 | 0.0 | A0A2C9WRV6_MANES; Uncharacterized protein | ||||
| STRING | cassava4.1_032088m | 0.0 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2118 | 30 | 86 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G45680.1 | 2e-88 | TCP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Manes.01G263300.1.p |



