 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MDP0000919694 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Malus
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 984aa MW: 109301 Da PI: 6.5689 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 130.2 | 7.5e-41 | 148 | 225 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C+adls+ak+yhrrhkvC +hska+++lv ++qrfCqqCsrfh+l+efDe+krsCrrrLa+hn+rrrk+++
MDP0000919694 148 VCQVEDCKADLSNAKDYHRRHKVCAMHSKATKALVGSVMQRFCQQCSRFHALQEFDEGKRSCRRRLAGHNRRRRKTHP 225
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 2.1E-32 | 143 | 210 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.932 | 146 | 223 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.16E-37 | 147 | 228 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 3.1E-29 | 149 | 222 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 1.2E-7 | 755 | 867 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 5.1E-6 | 757 | 863 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 8.33E-8 | 758 | 865 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 984 aa Download sequence Send to blast |
MESEFGGKAC NFRGVMVPNL KGVGKKSLEW DLNDFKWDGD LFTASPLNST PSDGRSRQLF 60 PARPETPSDA GLSNSSSSGS DNISPGNEKD QRELEKRRRD FFVENRELND EAASLNLKLG 120 GQTYPIMEEE VQTGKKTKTI GTTSNRAVCQ VEDCKADLSN AKDYHRRHKV CAMHSKATKA 180 LVGSVMQRFC QQCSRFHALQ EFDEGKRSCR RRLAGHNRRR RKTHPDTAVN GGSLNNEGGS 240 GYLLISLLRI LSNMHSSSSD QTKDQDVISH LLRSLANVAG TADGRNISTL LQGSQGLFNS 300 GTSVQTARRV LDMNAGVNTE DPLRSKGHCS ILPASRDSSE SKSVTPEATS RRFQLNDIDL 360 NSTYDDSQDY VENLGNSHVP ASPGTASLGF PSWMQRDSHK SSPPQTSGNS DLTSTQSPSS 420 SSGEAQSHTD RIIFKLFGKD PNELPLALRS QILDWLSHSP TNIESYIRPG CIILTIYLRL 480 EKSTWEEFCC HLGSSLKTLL DAADDPFWRT GWVYTRVQDY VAFTYNGEVV LDTPLPLKSN 540 KSCRISCIKP IAISLSERAE FIVKGFNLSS STTRLLCALE GKYLAQETCY DLLDDADSTV 600 EDDEQQCLKF SCSIPNVTGR GFIEVEDHGL SSSFFPFIVA EQEVCSEICM LEDVIEVSET 660 DDDIQSGPEK VEAKNQALDF IHELGWLLHR SRVKFRLGQL DPNLDTFPFR RFRLLMEFSI 720 DHDWCAVVKK LLGILFDGTV DAAEHPSLES ALLDMGLLHR AVRINSRRMV EFLLRFVPGL 780 TGSEQKEQVD RDGNSFLFKP DVVGPMGLTP LHIAASTDGC EQVLDALTDD PGKVGIKAWK 840 NTRDSTGLTP YDYACLRSRY SYVHIVQRKI SNTLESGHVV LDIPGLTLDR NGKQKQSDAH 900 KSSRVASLET ERNDIKAILR HCRLCEQKPA YSTTRSLVYR PAMLSLVAVA AVCVCVALLF 960 KSNPEVVFVL EPFRWEHLKF GSS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 2e-29 | 148 | 222 | 10 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mdo.8867 | 0.0 | bud| fruit | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed constitutively during plant development. {ECO:0000269|PubMed:10524240}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
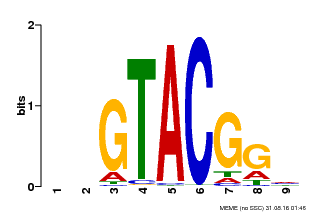
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009353416.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A498HJS4 | 0.0 | A0A498HJS4_MALDO; Uncharacterized protein | ||||
| STRING | XP_008390370.1 | 0.0 | (Malus domestica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1539 | 34 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | MDP0000919694 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



