 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MDP0000699611 | ||||||||
| Common Name | C3HL7 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Malus
|
||||||||
| Family | C3H | ||||||||
| Protein Properties | Length: 707aa MW: 77020.1 Da PI: 5.35 | ||||||||
| Description | C3H family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-CCCH | 21.2 | 5e-07 | 277 | 297 | 5 | 26 |
--SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 5 lCrffartGtCkyGdrCkFaHg 26
+C+ f++ GtC+ Gd C +aHg
MDP0000699611 277 PCPEFRK-GTCRQGDACEYAHG 297
8******.*************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF48403 | 1.34E-11 | 27 | 152 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 8.9E-15 | 29 | 155 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 1.1E-6 | 73 | 138 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 13.735 | 73 | 148 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.013 | 73 | 103 | IPR002110 | Ankyrin repeat |
| CDD | cd00204 | 2.69E-7 | 75 | 148 | No hit | No description |
| SMART | SM00248 | 0.64 | 108 | 140 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 9.938 | 108 | 143 | IPR002110 | Ankyrin repeat |
| SMART | SM00356 | 5.4E-5 | 272 | 298 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 1.3E-4 | 277 | 297 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 12.784 | 277 | 299 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 6.944 | 307 | 331 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 31 | 307 | 330 | IPR000571 | Zinc finger, CCCH-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 707 aa Download sequence Send to blast |
MCSGSKRNPS STDTIMEREK QEGTRFNFSI LLELAACDDX EGFKRAVEEE GLDVDEASYW 60 CGRLIGSKKL GFEXRTPLMV AAMFGSMNVL NYILQSCLVD VNKACGSDRA TALHCAVAGG 120 SAASAEVVKL LLAASADASS LDANGNQPGD LIAPAYSSSF GSRKKALEVM LKGVPSIDEP 180 FDFSEQMINE TEGQEQQEMT TPRASKDGTE KKEYPVDLSL PDIKNGIYST DEFRMYTFKV 240 KPCSRAYSHD WTECPFVHPG ENARRRDPRK YHYSCVPCPE FRKGTCRQGD ACEYAHGIFE 300 CWLHPAQYRT RLCKDETGCT RRVCFFAHKP EELRPLYAST GSAVPSPRSF SATAASLDMG 360 SITPLSLNSP SMMIPPDSTP PMTPTGPSSP MGGNMWQNNP NFAPPTLQLP GSRLKSTLSA 420 RDMDFEIEML SLERDRRXQQ RLIDEMSGSP SSWNKGLSPA SPFSASGNRT GELNTFGXVN 480 PTNLDDIFGS LDPAILPQFN GLSRDATASQ LHSPTGIQMR QNMNLQARPS YSASLSSSPV 540 RASPMFGVDA SSAAAVFNSR SAAFAKRSQS FIERSAGNRN SVVSSSADFG TIKPSNLSDW 600 GSPGGKLDWG IQGEELNKLR KSASFGFRSN GSSSPTASSM MPTNGDEPDV SWVQSLVKDG 660 PQASQQRGQF GFEDQQQQQC HPNNGGPEML PAWVEQLYFE QEQMVA* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mdo.2843 | 0.0 | bud| fruit| leaf| root| stem | ||||
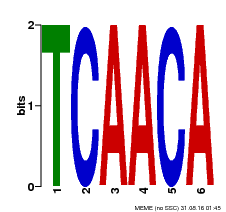
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00566 | DAP | Transfer from AT5G58620 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001315927.1 | 0.0 | zinc finger CCCH domain-containing protein 66-like | ||||
| Swissprot | Q9LUZ4 | 0.0 | C3H66_ARATH; Zinc finger CCCH domain-containing protein 66 | ||||
| TrEMBL | A0A498IRT1 | 0.0 | A0A498IRT1_MALDO; Uncharacterized protein | ||||
| STRING | XP_008384793.1 | 0.0 | (Malus domestica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1796 | 34 | 90 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G58620.1 | 0.0 | C3H family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | MDP0000699611 |



