 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MDP0000551046 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Malus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 602aa MW: 66916.6 Da PI: 6.594 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 20.7 | 1.1e-06 | 348 | 369 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
C +Cgk F+r nL+ H+r H
MDP0000551046 348 FCAICGKGFKRDANLRMHMRGH 369
6*******************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 4.16E-5 | 345 | 373 | No hit | No description |
| SMART | SM00355 | 0.012 | 347 | 369 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.884 | 347 | 374 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-5 | 348 | 398 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 349 | 369 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 53 | 396 | 429 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 21 | 434 | 456 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010044 | Biological Process | response to aluminum ion | ||||
| GO:0010447 | Biological Process | response to acidic pH | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 602 aa Download sequence Send to blast |
MHEFPRCLGC KKSLIKNXDT NKARRPGSVE KEASGQAIHV LVNECGMXXH GSCPMGIGGP 60 SSNIRYANFK CEDXSXEMDP KERQWDTWEN PSTGNDVTNT ISSDHQPFTS YKSRQHQQKW 120 ESPSVLDYEM GMELSFQKFH QPSDSQTSHT CNSNNDTKIP DREDGRMXEA QQPDKIQDWD 180 PRMTLNDLSF LEQKIHQLQD LVHVIVGRRG QVLGRPDELV AQQQQLITAD LTSIIAQLIS 240 TAGSLLPSVK HTLSTTIPXT GQFGQLGGSF IPSAAGNDVX VKLQVNSGSK LADQANHTDL 300 ISNYGTEHME EHETKDEEDA DEGENLPPGS YEILQLEKEE ILAPHTHFCA ICGKGFKRDA 360 NLRMHMRGHG DEYKTAAALA KPNKESSSEP TLIKRYSCPY AGCKRNKDHK KFQPLKTILC 420 VKNHYKRTHC DKSYTCSRCN SKKFSVIADL KTHEKHCGKD KWLCSCGTTF SRKDKLFGHI 480 ALFQGHTPAI PPDEPKGTLG PTNHGEGSEA SNRVGSINFS FGSTAPGGGG GSQNLMDVKE 540 SIDDPTSYFS PLNFETCNFD GFHELPRPPF EDSEISFLMP VSCNYTDKTG GESNFNNLQR 600 H* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mdo.8050 | 0.0 | bud| cell culture| fruit| leaf | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots (e.g. root tips and lateral roots), leaves, flowers (e.g. stigma, sepal, anther, and filament), stems, siliques and cotyledons. {ECO:0000269|PubMed:23935008}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. Together with STOP2, plays a critical role in tolerance to major stress factors in acid soils such as proton H(+) and aluminum ion Al(3+). Required for the expression of genes in response to acidic stress (e.g. ALMT1 and MATE), and Al-activated citrate exudation. {ECO:0000269|PubMed:17535918, ECO:0000269|PubMed:18826429, ECO:0000269|PubMed:19321711, ECO:0000269|PubMed:23935008}. | |||||
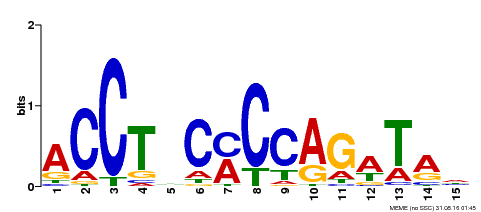
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00196 | ampDAP | Transfer from AT1G34370 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By shock H(+) and Al(3+) treatments. {ECO:0000269|PubMed:17535918}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008365851.3 | 0.0 | LOW QUALITY PROTEIN: protein SENSITIVE TO PROTON RHIZOTOXICITY 1-like | ||||
| Swissprot | Q9C8N5 | 0.0 | STOP1_ARATH; Protein SENSITIVE TO PROTON RHIZOTOXICITY 1 | ||||
| TrEMBL | A0A498J3B1 | 0.0 | A0A498J3B1_MALDO; Uncharacterized protein | ||||
| STRING | XP_008348808.1 | 0.0 | (Malus domestica) | ||||
| STRING | XP_008358329.1 | 0.0 | (Malus domestica) | ||||
| STRING | XP_008358333.1 | 0.0 | (Malus domestica) | ||||
| STRING | XP_008365851.1 | 0.0 | (Malus domestica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5340 | 32 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G34370.2 | 0.0 | C2H2 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | MDP0000551046 |



