 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MDP0000491277 | ||||||||
| Common Name | HD3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Malus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 826aa MW: 89687.4 Da PI: 6.4048 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 66.6 | 3.2e-21 | 120 | 175 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+++l+L++rqVk+WFqNrR+++k
MDP0000491277 120 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSRRLNLETRQVKFWFQNRRTQMK 175
688999***********************************************999 PP
| |||||||
| 2 | START | 205.2 | 2.6e-64 | 327 | 550 | 1 | 205 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEE CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetle 83
ela++a++elvk+a+ +ep+W +s e +n +e++++f++ + + +ea+r+sg v+ ++ +lve+l+d++ +W e+++ + +t++
MDP0000491277 327 ELALAAMDELVKMAQTDEPLWLRSLeggrEVLNHEEYMRNFTPCIGlkpngFVSEASRESGTVIINSLTLVETLMDSN-RWLEMFPgvlaRTSTTD 421
5899************************9*************99989*******************************.***************** PP
EECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE- CS
START 84 vissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvdl 172
vissg galqlm aelq+lsplvp R++ f+R+++q+ +g+w++vdvSvd ++ + + ++ +++lpSg+++++++ng+skvtwveh+++
MDP0000491277 422 VISSGmggtrnGALQLMHAELQVLSPLVPvREVNFLRFCKQHAEGVWAVVDVSVDAIRDTTGVPTFMNCRRLPSGCVVQDMPNGYSKVTWVEHAEY 517
***********************************************************999********************************** PP
-SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 173 kgrlphwllrslvksglaegaktwvatlqrqce 205
+++++h l+r+l++sg+ +ga++wvatlqrq e
MDP0000491277 518 DESQVHHLYRPLLSSGMGFGAQRWVATLQRQSE 550
******************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.45E-21 | 101 | 177 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.7E-22 | 105 | 177 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 18.058 | 117 | 177 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 3.6E-19 | 118 | 181 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.19E-19 | 120 | 177 | No hit | No description |
| Pfam | PF00046 | 8.1E-19 | 120 | 175 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 152 | 175 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 42.295 | 318 | 554 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 8.63E-33 | 319 | 550 | No hit | No description |
| CDD | cd08875 | 2.67E-123 | 322 | 550 | No hit | No description |
| SMART | SM00234 | 1.1E-47 | 327 | 551 | IPR002913 | START domain |
| Pfam | PF01852 | 4.7E-56 | 327 | 550 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 4.3E-22 | 579 | 748 | No hit | No description |
| SuperFamily | SSF55961 | 4.3E-22 | 778 | 818 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 826 aa Download sequence Send to blast |
MSFGGFLDNS TGSGVGARIV ADIPYTNNSN NMPSSAIAQP RLVTQSLTKS MFNSPGLSLA 60 LQTNVDGQGD VTRVAENYEA NNGGRRSREE EHESRSGSDN MDGASGDDQD AADNHHPRKK 120 KRYHRHTPQQ IQELEALFKE CPHPDEKQRL ELSRRLNLET RQVKFWFQNR RTQMKTQLER 180 HENSLLRQEN DKLRAENMSI REAMRNPICS NCGGPAIIGD ISLEEQHLRI ENARLKDDLD 240 RVCALAGKFL GRPISSLAAS MGPPLPSSTL ELGVGSNGFG GMSNVATSMS MGNDFGGGIG 300 SAMSVVSHGR PSVTGLDRSM ERSIFLELAL AAMDELVKMA QTDEPLWLRS LEGGREVLNH 360 EEYMRNFTPC IGLKPNGFVS EASRESGTVI INSLTLVETL MDSNRWLEMF PGVLARTSTT 420 DVISSGMGGT RNGALQLMHA ELQVLSPLVP VREVNFLRFC KQHAEGVWAV VDVSVDAIRD 480 TTGVPTFMNC RRLPSGCVVQ DMPNGYSKVT WVEHAEYDES QVHHLYRPLL SSGMGFGAQR 540 WVATLQRQSE CQAILMSSCV TSRDHTAITA SGRRSMLKLA QRMTDNFCAG VCASTVHKWT 600 KLNAGNVDED VRVMTRESLD DPGEPPGVVL SAATSVWLPV SPQRLFDFLR DERLRSEWDI 660 LSNGGPMQEM AHIAKGQDPG NCVSLLRARA NANQGSMLIL QETCIDAAGS LVVYAPVDIP 720 AMHVVMNGGD SAYVALLPSG FAIVPDGPGS RGPMFGKGGS HGSGNSGGGV DDGGHRVSGS 780 LLTMTFQILV NSLPTAKLTV ESVETVNHLI SCTVQKIKAA LHCES* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mdo.2928 | 0.0 | bud| fruit| leaf| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves and floral buds. {ECO:0000269|PubMed:10402424}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
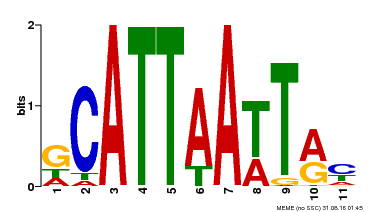
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF067961 | 0.0 | AF067961.1 Malus domestica homeodomain protein (h3) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008385703.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X1 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A498JIK4 | 0.0 | A0A498JIK4_MALDO; Uncharacterized protein | ||||
| STRING | XP_008385703.1 | 0.0 | (Malus domestica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1192 | 34 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G61150.1 | 0.0 | homeodomain GLABROUS 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | MDP0000491277 |
| Entrez Gene | 103445972 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



