 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MDP0000325672 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Malus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 353aa MW: 38711.6 Da PI: 9.5385 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 42.2 | 1.9e-13 | 81 | 125 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT++E++++++a +++ + Wk+I + +g +t q++s+ qky
MDP0000325672 81 RESWTEQEHDKFLEALQLFDRD-WKKIEAFIG-SKTVIQIRSHAQKY 125
789*****************77.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.32E-16 | 75 | 131 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 14.986 | 76 | 130 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 8.1E-7 | 79 | 131 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.5E-18 | 79 | 128 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 7.9E-11 | 80 | 128 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.9E-11 | 81 | 125 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.55E-8 | 83 | 126 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 353 aa Download sequence Send to blast |
MLSLNPNPAQ GFYLFDPMNT NMGLPGLNPM PPPTTGAASA TAATSTTAAC TSSSAPANAA 60 SMSADDQSKK IRKPYTITKS RESWTEQEHD KFLEALQLFD RDWKKIEAFI GSKTVIQIRS 120 HAQKYFLKVQ KKGTSEHVPP PRPKRKATHP YPQKAPKSAA XVSQVAGPFQ SSSXLLEHGY 180 VYQPDSSIVL GTPVNAATLL SWSYNSVPPV NVXQTIKDEG RLSGPTVTHN SCYSSSNESN 240 PTNWKVSETV DRVDPGQPHR VLPDFAQVYK FIGSVFDPSA ISHLERLRQL DPINFETALL 300 LMRNLSINLT SPEFQDHRKL ITSYAVGSDK TKSDGNCNFS RIGKSENAIL SAX |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mdo.16616 | 0.0 | bud| fruit| leaf | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
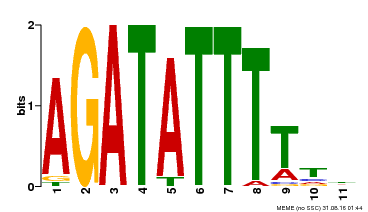
| Motif ID | Method | Source | Motif file |
| MP00423 | DAP | Transfer from AT4G01280 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008354011.2 | 0.0 | protein REVEILLE 3-like | ||||
| Swissprot | C0SVG5 | 1e-102 | RVE5_ARATH; Protein REVEILLE 5 | ||||
| Swissprot | Q6R0H0 | 1e-103 | RVE3_ARATH; Protein REVEILLE 3 | ||||
| Swissprot | Q8H0W3 | 1e-102 | RVE6_ARATH; Protein REVEILLE 6 | ||||
| TrEMBL | A0A498KI09 | 0.0 | A0A498KI09_MALDO; Uncharacterized protein | ||||
| STRING | XP_008343412.1 | 0.0 | (Malus domestica) | ||||
| STRING | XP_008345636.1 | 0.0 | (Malus domestica) | ||||
| STRING | XP_008354011.1 | 0.0 | (Malus domestica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1119 | 33 | 96 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G52660.1 | 1e-101 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | MDP0000325672 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



