 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MDP0000313059 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Malus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 914aa MW: 100377 Da PI: 6.9183 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.5 | 2.3e-18 | 17 | 75 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++ +++ps +r++L +++ +++ +q+kvWFqNrR +ek+
MDP0000313059 17 KYVRYTPEQVEALERLYHDCPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 75
5679*****************************************************97 PP
| |||||||
| 2 | START | 164.9 | 5.8e-52 | 161 | 367 | 2 | 203 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEEEEEE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..galqlmv 95
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+l+d + W ++++++++l+v+ ++ g+++l +
MDP0000313059 161 IAEETLAEFLSKATGTAVEWVQMPGMKPGPDSIGIVAISHGCTGVAARACGLVGLEPT-RVAEILKDLPSWLRDCRAVDVLNVLPTAngGTIELLY 255
7899******************************************************.**************************9999******* PP
EXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHH CS
START 96 aelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslvks 187
++l+a+++l+p df+ +Ry+ l++g++v++ +S+ ++q+ p+ +++vRae+lpSg+li+p+++g+s +++v+h+dl+ +++++lr+l++s
MDP0000313059 256 MQLYAPTTLAPaCDFWLLRYTSVLEDGSLVVCARSLKNTQNGPTmppVQHFVRAEMLPSGYLIRPCEGGGSIIHIVDHMDLEPCSVPEVLRPLYES 351
******************************************9998899*********************************************** PP
HHHHHHHHHHHHTXXX CS
START 188 glaegaktwvatlqrq 203
+++ ++k+++a +r
MDP0000313059 352 SAVLAQKMTMAGSKRT 367
***********98875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.386 | 12 | 76 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-15 | 14 | 80 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 9.84E-17 | 16 | 80 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.51E-16 | 17 | 77 | No hit | No description |
| Pfam | PF00046 | 6.5E-16 | 18 | 75 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 3.9E-18 | 19 | 75 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.90E-6 | 69 | 108 | No hit | No description |
| PROSITE profile | PS50848 | 22.545 | 151 | 352 | IPR002913 | START domain |
| CDD | cd08875 | 5.78E-73 | 155 | 362 | No hit | No description |
| SuperFamily | SSF55961 | 1.21E-33 | 160 | 353 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 7.2E-19 | 160 | 342 | IPR023393 | START-like domain |
| SMART | SM00234 | 1.5E-38 | 160 | 370 | IPR002913 | START domain |
| Pfam | PF01852 | 1.6E-49 | 161 | 367 | IPR002913 | START domain |
| Pfam | PF08670 | 1.2E-49 | 728 | 867 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 914 aa Download sequence Send to blast |
MAMSCKDGNK HALDNGKYVR YTPEQVEALE RLYHDCPKPS SIRRQQLIRE CPILSNIEPK 60 QIKVWFQNRR CREKQRKEAS RLQAVNRKLS AMNKLLMEEN DRLQKQVSHL VYENGYFRQH 120 TQGTTLATKD TSCESVVTSG QHHLTPQHPP RDASPAGLLS IAEETLAEFL SKATGTAVEW 180 VQMPGMKPGP DSIGIVAISH GCTGVAARAC GLVGLEPTRV AEILKDLPSW LRDCRAVDVL 240 NVLPTANGGT IELLYMQLYA PTTLAPACDF WLLRYTSVLE DGSLVVCARS LKNTQNGPTM 300 PPVQHFVRAE MLPSGYLIRP CEGGGSIIHI VDHMDLEPCS VPEVLRPLYE SSAVLAQKMT 360 MAGSKRTKLH VNLSXKHAVK ALRQLRQIAH EVSQSNVTGW GRRPAALRAL SQRLSREGFF 420 VSWKLMEFSA RGFNDALNGF TDEGWSMMGN DGMDDVTILI NSSPDKLMGL NLSFGNGFPA 480 VSNSVLCAKA SMLLQNVPPA ILLRFLREHR SEWADNNIDA YSAAAVKVGP CSLAGSRVGS 540 FGXQVILPLA HTLEHEEFLE VIKLEGVGHS PEDAMMPREM FLLQLCSGMD ENAVGSCAEL 600 IFAPIDASFA DDAPLLPSGF RIIPLDYGKE ASSPNRTLDL ASALEIGPTG NKGSSEYSAS 660 AGCVRSVMTI AFEFACETHM QEHVASMARQ YVRSIISSVQ RVALALSPSN LSSQAGLRSP 720 LGTPEAQTLA RWICNSYRCY LGVELLKSGN EGSESILKSL WHHSDAIMCC SLKALPVFTF 780 ANQAGLDMLE TTLVALQDIT LEKIFDDHGR KTLCSEFPQI MQQGFTCLQG GICLSSMGRP 840 VSYERAVAWK VLNEEETAHC MCFLFVNCYN NGGWGAILAD VYARKEKEVS QVINGTAFIQ 900 TNLVEALKTV QLKP |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mdo.12971 | 1e-179 | bud| leaf| root| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed the developing vascular elements and the adaxial portion of cotyledons. Expressed in developing ovules, stamens and carpels. Expressed in procambium and shoot meristem. {ECO:0000269|PubMed:14701930, ECO:0000269|PubMed:15705957}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
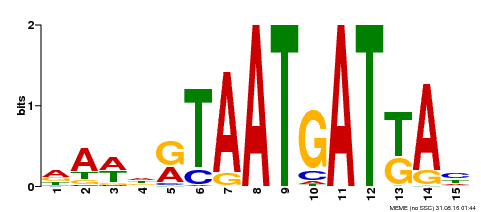
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FJ177426 | 0.0 | FJ177426.1 Malus x domestica putative HB15 HD-ZipIII mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001280898.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Refseq | XP_009353561.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | B6DXL7 | 0.0 | B6DXL7_MALDO; Putative HB15 HD-ZipIII | ||||
| STRING | XP_009353561.1 | 0.0 | (Pyrus x bretschneideri) | ||||
| STRING | XP_008380947.1 | 0.0 | (Malus domestica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1946 | 33 | 87 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | MDP0000313059 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



