 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MDP0000298233 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Malus
|
||||||||
| Family | C3H | ||||||||
| Protein Properties | Length: 573aa MW: 64288.5 Da PI: 8.9956 | ||||||||
| Description | C3H family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-CCCH | 30 | 9.2e-10 | 37 | 61 | 3 | 27 |
S---SGGGGTS--TTTTT-SS-SSS CS
zf-CCCH 3 telCrffartGtCkyGdrCkFaHgp 27
+ +C++++r+G CkyG C+++H++
MDP0000298233 37 QTECKYYLRSGGCKYGKDCRYSHSK 61
568********************95 PP
| |||||||
| 2 | zf-CCCH | 36.5 | 8.2e-12 | 81 | 106 | 1 | 26 |
--S---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 1 yktelCrffartGtCkyGdrCkFaHg 26
+++++C++++rtG Cky ++C+F+H+
MDP0000298233 81 PGERECPYYMRTGSCKYASNCRFNHP 106
5789*********************9 PP
| |||||||
| 3 | zf-CCCH | 26.6 | 1.1e-08 | 217 | 243 | 1 | 27 |
--S---SGGGGTS--TTTTT-SS-SSS CS
zf-CCCH 1 yktelCrffartGtCkyGdrCkFaHgp 27
++++ C++f+rtG Ck+ ++Ck++H++
MDP0000298233 217 PGQPDCSYFLRTGDCKFKSNCKYHHPK 243
5789*********************96 PP
| |||||||
| 4 | zf-CCCH | 33.3 | 8.2e-11 | 265 | 288 | 3 | 26 |
S---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 3 telCrffartGtCkyGdrCkFaHg 26
+ +C +++r+G+Ck+G+ CkF H+
MDP0000298233 265 QNICTHYSRYGICKFGPACKFDHS 288
679********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50103 | 15.591 | 34 | 62 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 1.4E-5 | 34 | 61 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 1.14E-5 | 37 | 61 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 2.5E-7 | 38 | 61 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 3.5E-5 | 40 | 60 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 15.131 | 80 | 108 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 7.5E-6 | 80 | 107 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 1.1E-8 | 82 | 106 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 1.31E-5 | 83 | 107 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 1.1E-4 | 86 | 105 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 14.352 | 216 | 244 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 1.3E-4 | 216 | 243 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 3.5E-6 | 218 | 243 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 6.67E-5 | 221 | 244 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 4.45E-6 | 260 | 288 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 14.446 | 262 | 290 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 2.6E-4 | 262 | 289 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 4.1E-8 | 265 | 288 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 0.001 | 267 | 289 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:3.40.50.1000 | 0.001 | 532 | 569 | IPR023214 | HAD-like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 573 aa Download sequence Send to blast |
MENLMSCRSW IRQRMWRRTE LVPRERVKDE SAENPSQTEC KYYLRSGGCK YGKDCRYSHS 60 KVKPSVAPVL ELNFLGLPIR PGERECPYYM RTGSCKYASN CRFNHPDPTA IGGSDPPSXX 120 GKDGPASLQV ASQXTVAXWS APRPLNEAPV YTPMMIPPPQ GVSSQNSEWN GCRAPAYIPE 180 SSMPAXPPYM MNNSVTETNV YKQYPLQXQV EEFPERPGQP DCSYFLRTGD CKFKSNCKYH 240 HPKTQTAVAP LFTLSDKGLP LRPDQNICTH YSRYGICKFG PACKFDHSSH ISSSTTSGPD 300 HQLPYGDWAT TNGAGIAGSR SGTDASVVYR TCCGVCVVCC MSWHRRGPVK VTVNSXHISQ 360 RISXLRSRHR EGMYEVDGEL ISYEXTYTRT KFNGIKWDSL SLTPSLGGPS ETPKSERTAK 420 TTSLSRRLAR LLALLDRRQR TSSLSPVQPN FYSRLPSQFS LLPACPKRTP WPPSLFCFVS 480 VTKFTWEGVR KLSNDSRSTL NLLHFLPNRS WDRLLVVVDK SPECGDGGAE GYWIRKCEDA 540 WHLCDPDAEK VVEALRKAGV KVAVVSNFDA RLX |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mdo.13055 | 1e-119 | fruit| leaf| root | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00582 | DAP | Transfer from AT5G63260 | Download |
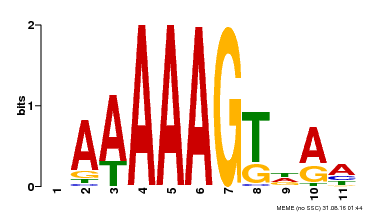
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028959756.1 | 0.0 | zinc finger CCCH domain-containing protein 67-like | ||||
| TrEMBL | A0A498JJL4 | 0.0 | A0A498JJL4_MALDO; Uncharacterized protein | ||||
| STRING | XP_008337071.1 | 0.0 | (Malus domestica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4156 | 33 | 61 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G63260.1 | 1e-102 | C3H family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | MDP0000298233 |



