 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MDP0000291729 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Malus
|
||||||||
| Family | YABBY | ||||||||
| Protein Properties | Length: 182aa MW: 20182.9 Da PI: 8.2617 | ||||||||
| Description | YABBY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | YABBY | 170.8 | 9.3e-53 | 13 | 167 | 4 | 163 |
YABBY 4 fssseqvCyvqCnfCntilavsvPstslfkvvtvrCGhCtsllsvnlakasqllaaeshldeslkeelleelkveeenlksnvekeesastsvsse 99
++ se++Cyv+CnfCnt+lav +P + l+ +vtv+CGhC++l ++ + q + +h +sl+ + ++ +e+++ n+ k++++sts s
MDP0000291729 13 NQPSEHLCYVRCNFCNTVLAVGIPCKRLMDTVTVKCGHCSNLSFLSTRPPLQGQCLPDH-PTSLTLQESDQYWNEKQAGCCNDFKKDQSSTSSS-- 105
5789*************************************988888888888888887.3344444444444555555556666555555444.. PP
YABBY 100 klsenedeevprvppvirPPekrqrvPsaynrfikeeiqrikasnPdishreafsaaaknWahf 163
+ ++d +p++p v++PPek+ r Psaynrf+k+eiqrika+nP+i hreafsaaaknWa +
MDP0000291729 106 --PVSSDPPSPKAPFVVKPPEKKHRLPSAYNRFMKDEIQRIKAANPEIPHREAFSAAAKNWARY 167
..35678899****************************************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF04690 | 3.3E-56 | 15 | 167 | IPR006780 | YABBY protein |
| SuperFamily | SSF47095 | 9.03E-9 | 111 | 165 | IPR009071 | High mobility group box domain |
| Gene3D | G3DSA:1.10.30.10 | 5.6E-5 | 121 | 166 | IPR009071 | High mobility group box domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010254 | Biological Process | nectary development | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0048479 | Biological Process | style development | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 182 aa Download sequence Send to blast |
MNMEEKVTMD LINQPSEHLC YVRCNFCNTV LAVGIPCKRL MDTVTVKCGH CSNLSFLSTR 60 PPLQGQCLPD HPTSLTLQES DQYWNEKQAG CCNDFKKDQS STSSSPVSSD PPSPKAPFVV 120 KPPEKKHRLP SAYNRFMKDE IQRIKAANPE IPHREAFSAA AKNWARYIPC QSSAAGSGSG 180 SY |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Detected during flower and leaf development. Expression in the flower meristem in the early stage of flower development. When carpel primordia begin to form, specific and uniform expression in carpel primordia. Expression in the central region of the leaf plastochron 1 (P1) primordia. Detected up to P4 stage, hardly detected in the P5 leaves. {ECO:0000269|PubMed:14729915}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Regulates carpel specification in flower development. Severe or intermediate mutation in DL causes complete or partial homeotic conversion of carpels into stamens without affecting the identities of other floral organs. Interacts antagonistically with class B genes and controls floral meristem determinacy. Regulates midrib formation in leaves probably by inducing cell proliferation in the central region of the leaf. {ECO:0000269|PubMed:14729915}. | |||||
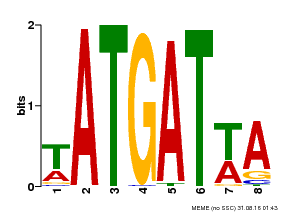
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00220 | DAP | Transfer from AT1G69180 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB627250 | 1e-137 | AB627250.1 Malus x domestica mRNA, microsatellite: MEST098, clone: FLW01_10_E08. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008339784.1 | 1e-104 | protein CRABS CLAW isoform X1 | ||||
| Swissprot | Q76EJ0 | 2e-64 | YABDL_ORYSJ; Protein DROOPING LEAF | ||||
| TrEMBL | A0A498IBD4 | 1e-102 | A0A498IBD4_MALDO; Uncharacterized protein | ||||
| STRING | XP_008339784.1 | 1e-103 | (Malus domestica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF10259 | 30 | 40 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69180.1 | 3e-64 | YABBY family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | MDP0000291729 |



