 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MDP0000272003 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Malus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 328aa MW: 36937.9 Da PI: 8.7487 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100.7 | 9.9e-32 | 69 | 122 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH rF++ave+LGG+++AtPk +l+lm++kgL ++hvkSHLQ+YR+
MDP0000272003 69 PRLRWTPDLHLRFIHAVERLGGQDRATPKLVLQLMNIKGLNIAHVKSHLQMYRS 122
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 13.002 | 65 | 125 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.1E-30 | 65 | 123 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.51E-14 | 67 | 123 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.5E-22 | 69 | 124 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 8.9E-9 | 70 | 121 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 328 aa Download sequence Send to blast |
MMGSQGGSEC SKTSPSDKQN EDGSESGENY SNGENSKPKN NGGSSSNSTV EESDQKKTSV 60 RPYVRSKMPR LRWTPDLHLR FIHAVERLGG QDRATPKLVL QLMNIKGLNI AHVKSHLQMY 120 RSKKIDDAGQ VLADHQGHLV ECGDKNIYNL SQLPMLQGYN RSHTTSFRYG SYGDTNSWTA 180 LENPRFSRSS TGFQGNWSTS STGNNAYNYL TSCSFGDHPQ SSWITRTLKE ECQLHNIRES 240 LKPKEFITSF NTNRGPFRDS DTISKNLVQD QSKMLKRKAS DCDDLDLDLS LRLTSKKNDE 300 DHTHEVDSNL SLCLYSPPTS SKLRRLKE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 3e-17 | 70 | 124 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 3e-17 | 70 | 124 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 3e-17 | 70 | 124 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 3e-17 | 70 | 124 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
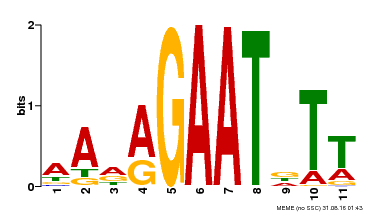
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008363851.1 | 0.0 | two-component response regulator ARR18-like | ||||
| TrEMBL | A0A498K210 | 0.0 | A0A498K210_MALDO; Uncharacterized protein | ||||
| STRING | XP_008363851.1 | 0.0 | (Malus domestica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2378 | 34 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 1e-59 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | MDP0000272003 |



