 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSMUA_Achr8P02380_001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Zingiberales; Musaceae; Musa
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 299aa MW: 33228.8 Da PI: 8.4564 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 34.5 | 4.7e-11 | 37 | 85 | 2 | 48 |
SSS-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 2 grWTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrwqkyl 48
g+WT+eE +++ a +++ ++ +W+ +a+ ++ g+t+ ++ s+++++l
GSMUA_Achr8P02380_001 37 GNWTPEENKRFEYALAKFDKDtpdRWEQVAASIP-GKTAWDVESHYRDLL 85
89********************************.************986 PP
| |||||||
| 2 | Myb_DNA-binding | 42.3 | 1.7e-13 | 144 | 188 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT++E+++++ + k++G+g+W+ I+r + +Rt+ q+ s+ qky
GSMUA_Achr8P02380_001 144 PWTEDEHKRFLFGLKKYGKGDWRNISRNFVITRTPTQVASHAQKY 188
8*****************************99************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 7.329 | 34 | 87 | IPR017884 | SANT domain |
| SMART | SM00717 | 6.9E-11 | 35 | 87 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 1.51E-12 | 37 | 91 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-6 | 37 | 86 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 6.5E-9 | 37 | 85 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.34E-8 | 38 | 85 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 3.9E-11 | 135 | 189 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 18.121 | 137 | 193 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 6.75E-17 | 139 | 193 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.3E-17 | 140 | 191 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.2E-10 | 141 | 191 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 7.83E-9 | 144 | 189 | No hit | No description |
| Pfam | PF00249 | 1.2E-11 | 144 | 188 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 299 aa Download sequence Send to blast |
MTTRSWMEVL PPATVPCYPS SSWFIGEKMS GGGIGGGNWT PEENKRFEYA LAKFDKDTPD 60 RWEQVAASIP GKTAWDVESH YRDLLDDVSD IEAGRIPCPG YDSSSFTLDW ETNYGFEAST 120 QPYCIGGKRS AARASDQERK KGVPWTEDEH KRFLFGLKKY GKGDWRNISR NFVITRTPTQ 180 VASHAQKYFI RLNSGGKDKR RSSIHDITTA NLPDNRPPSP SQSSDPATQT SLASTPLPSA 240 PFSSILDSSH PNEATTIATS SVQGSTIRAT KLWSDTLWSA TRRSCTSEWH TRCYRGSGP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 7e-17 | 36 | 104 | 8 | 76 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the dorsovental asymmetry of flowers. Promotes ventral identity. {ECO:0000269|PubMed:11937495, ECO:0000269|PubMed:9118809}. | |||||
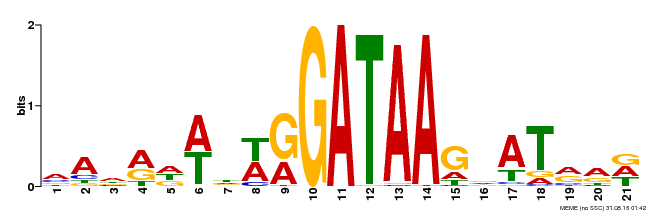
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00296 | DAP | Transfer from AT2G38090 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009402284.1 | 1e-134 | PREDICTED: transcription factor DIVARICATA-like | ||||
| Swissprot | Q8S9H7 | 3e-89 | DIV_ANTMA; Transcription factor DIVARICATA | ||||
| TrEMBL | M0TLZ0 | 0.0 | M0TLZ0_MUSAM; Uncharacterized protein | ||||
| STRING | GSMUA_Achr8P02380_001 | 0.0 | (Musa acuminata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2274 | 37 | 96 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38090.1 | 2e-88 | MYB family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



