 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSMUA_Achr1P06360_001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Zingiberales; Musaceae; Musa
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 726aa MW: 79534.7 Da PI: 8.5948 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 49.4 | 1e-15 | 25 | 69 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT+eE+ ++++a k++G W +I ++g ++t+ q++s+ qk+
GSMUA_Achr1P06360_001 25 REKWTEEEHNRFLEALKLYGRA-WQRIEDHIG-TKTAVQIRSHAQKF 69
789*****************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.59E-16 | 19 | 75 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 20.234 | 20 | 74 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.4E-16 | 23 | 72 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.5E-12 | 24 | 72 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-8 | 25 | 65 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 3.2E-13 | 25 | 68 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.85E-9 | 27 | 70 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 726 aa Download sequence Send to blast |
MEGNSSGDEM VVVKTRKPYT ITKQREKWTE EEHNRFLEAL KLYGRAWQRI EDHIGTKTAV 60 QIRSHAQKFF TKLEKEALEK GIPIGQSHAI DIPPPRPKRK PSSPYPRKSS TCCLTPGETI 120 NGKSSRSMFL GANKVVDIKS GAPQEKFTAT QKLQNEEISE PSQSEVLNVF QDALSTSISS 180 VNKSSSNHSK YMEILPTDEK VDDKIASSKS SAPHEVDKEL KKNGKAFIGQ ETEGVQGNCT 240 KPHINLALDK GESASKLHGL TSKDSVKGDQ TQPRHTMGRN GARNMHTEDS DGQNPTSVTG 300 QVEGHANDIS SMNPEASAIP VQNISRVNSM HHPFPAFAPF THFHSSQDAY RSFLNISSTF 360 SSFILSTLLQ NPAVHAAARL AASSCPSAEV EASLQSTPVF MAGERHINPA PGLEAIAAVT 420 VAAAAAWWKA HGLLPWFPPA AFAFSPPNTT TIPSADTAQL HSHGCTLEKP LAEDQQIGKQ 480 NLYEDLKPPR HSSKSLSLSL SSSDSHDSGR GKNSSELEDS TSNKFKPLVA SGFDDSDNPR 540 NKKKQDRSSC GSNTPSSSEV ETDNAGKHEK VNDEAKEAYF NNSAACETNQ CRFRSSGHMN 600 ESWKAVSEDG RLAFQQLFKR EVLPQSFPPP LDIAAATTIK KGETSKLLVD LNKNICSATD 660 FNHLRGHTKE EVCIRSKDRI THGKLKFHQT GFKPYKRCSV EAKENRAAAE ENNGKKKMRL 720 QGEAST |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
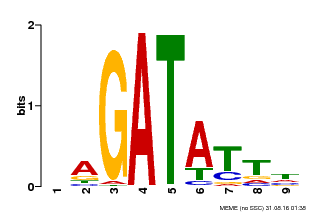
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009395074.1 | 0.0 | PREDICTED: protein LHY | ||||
| TrEMBL | M0RXJ4 | 0.0 | M0RXJ4_MUSAM; Uncharacterized protein | ||||
| STRING | GSMUA_Achr1P06360_001 | 0.0 | (Musa acuminata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6031 | 34 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01060.4 | 4e-40 | MYB_related family protein | ||||



