 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lus10032541 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Linaceae; Linum
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 850aa MW: 94300.5 Da PI: 6.7756 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 70.2 | 2.7e-22 | 161 | 262 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE.. CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefel 92
f+k+lt sd++++g +++ +++a+e+ + + + +++l+ +d+++ +W++++i+r++++r++l++GW+ Fv++++L +gD ++F + +++el
Lus10032541 161 FCKTLTASDTSTHGGFSVLRRHADEClppldmSRQ-PPTQELVAKDLHSIEWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL--RGENGEL 255
99*****************************7333.34459************************************************..449999* PP
EEEEE-S CS
B3 93 vvkvfrk 99
+v+v+r+
Lus10032541 256 RVGVRRA 262
****996 PP
| |||||||
| 2 | Auxin_resp | 117.3 | 1.3e-38 | 287 | 369 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha+st++ F+v+Y+Pr+s+s+F+v++++++++ k+++++GmRfkm+fe+e+++e+r++Gt+vg++d+d+ rW++SkWr+Lk
Lus10032541 287 AWHAISTGTLFTVYYKPRTSPSQFIVPYDQYMESAKSNYTIGMRFKMRFEGEEAPEQRFTGTIVGIEDADSKRWQESKWRCLK 369
89*******************************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 3.01E-40 | 157 | 290 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 6.0E-40 | 158 | 276 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.83E-20 | 159 | 261 | No hit | No description |
| Pfam | PF02362 | 4.1E-20 | 161 | 262 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.237 | 161 | 263 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.4E-22 | 161 | 263 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 3.4E-36 | 287 | 369 | IPR010525 | Auxin response factor |
| SuperFamily | SSF54277 | 2.4E-12 | 723 | 802 | No hit | No description |
| PROSITE profile | PS51745 | 25.683 | 725 | 809 | IPR000270 | PB1 domain |
| Pfam | PF02309 | 1.2E-7 | 770 | 810 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 850 aa Download sequence Send to blast |
MASSEISMKG NSGNVRGDSF SSSYSEQTEG RKAIEGQKGV PLAHPGAPID AEAALYKELW 60 HACAGPLVTV PRKGELVYYF PQGHIEQVEA STNQGADQQM PLYNLPSKIL CRVLNVELKA 120 EPDTDEVFAQ VTLLPEANQD ENAVKKEPPL PPPPRFHVHS FCKTLTASDT STHGGFSVLR 180 RHADECLPPL DMSRQPPTQE LVAKDLHSIE WRFRHIFRGQ PRRHLLQSGW SVFVSSKRLV 240 AGDAFIFLRG ENGELRVGVR RAMRQQGNVP SSVISSHSMH LGVLATAWHA ISTGTLFTVY 300 YKPRTSPSQF IVPYDQYMES AKSNYTIGMR FKMRFEGEEA PEQRFTGTIV GIEDADSKRW 360 QESKWRCLKV RWDENTSIPR PERVSPWQIE PALAPPALNP LPMARPKRPR TSIIPSSPDS 420 SVLTREVLSK VTADPAPASG YSRVLQGQEL STLRGNFAES TESDTAEKSV RWPLSVDDEK 480 IDVVSADRRT GHESWMPPVR HEPTYTDLLS GFAGNADSSS HGYSSPFADR GSFHGNSMRK 540 RGVDQDGKFS LLGNPWSLMP SGLSLKVSDS NVKAPWQSGD LGTPVRANSR YSAFSEYARA 600 DQSHINWLMP PPSQFDHHLQ TRDMMSKSIL APDRDAGKST DGNCKLFGIP LISNSKVADV 660 AASHKSTMND SVDHTNASSQ HHSREYDQLE QSKSTKMNDD SEHENRVEHS HVRDNHGKSQ 720 CGSTRSCTKV HKQGIALGRS VDLSKFNNYD QLIAELDRLF EFGGELIDPK QNWMIVYTDD 780 EGDMMLVGDD PWQEFVGMVR KIFIYTREEV QKMNPGSLNA KGNSNCDGEG GEMKQSAAAC 840 SLASASENC* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 0.0 | 54 | 391 | 18 | 354 | Auxin response factor 1 |
| 4ldw_A | 0.0 | 54 | 391 | 18 | 354 | Auxin response factor 1 |
| 4ldw_B | 0.0 | 54 | 391 | 18 | 354 | Auxin response factor 1 |
| 4ldx_A | 0.0 | 54 | 391 | 18 | 354 | Auxin response factor 1 |
| 4ldx_B | 0.0 | 54 | 391 | 18 | 354 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 405 | 411 | PKRPRTS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Could act as transcriptional activator or repressor. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Promotes flowering, stamen development, floral organ abscission and fruit dehiscence. Functions independently of ethylene and cytokinin response pathways. May act as a repressor of cell division and organ growth. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:15960614, ECO:0000269|PubMed:16176952, ECO:0000269|PubMed:16339187}. | |||||
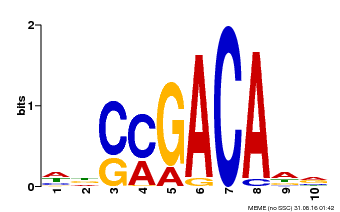
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lus10032541 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012090599.1 | 0.0 | auxin response factor 2 | ||||
| Swissprot | Q94JM3 | 0.0 | ARFB_ARATH; Auxin response factor 2 | ||||
| TrEMBL | A0A067JF52 | 0.0 | A0A067JF52_JATCU; Auxin response factor | ||||
| STRING | Lus10032541 | 0.0 | (Linum usitatissimum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3281 | 33 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Lus10032541 |



