 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lus10021699 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Linaceae; Linum
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 357aa MW: 39034.4 Da PI: 6.9047 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100 | 1.5e-31 | 215 | 270 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k r++W+peLH+rF++a++qLGGs++AtPk+i+elmkv+gLt+++vkSHLQk+Rl+
Lus10021699 215 KRRRCWSPELHRRFLHALQQLGGSHVATPKQIRELMKVDGLTNDEVKSHLQKFRLH 270
689***************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.704 | 212 | 272 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-27 | 212 | 273 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 4.41E-18 | 213 | 273 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.4E-26 | 215 | 270 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.1E-6 | 217 | 267 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 357 aa Download sequence Send to blast |
MDSLHHKKRS NEYAGALEEE RRKIIMFQRE LPLSLELVTQ AIEACRRDLS GTTTEHEKSD 60 CSEQTSGECP VTEEFIPIIK ANSSSCDDED DNYEKKRPLK RTRSKSDWLR SVQLWNQPPD 120 PPSPKQAVPR KALVTEVKRN GAFQPFQKEK SNGDDNITVA PAAVVSAPPS KVLLPSAMSS 180 TADATPTAAG PSSVVSDSEN KQKEDEDKES QAKRKRRRCW SPELHRRFLH ALQQLGGSHV 240 ATPKQIRELM KVDGLTNDEV KSHLQKFRLH TRRPSHSVQN GSNSAGGPQF VVVGGIWVPP 300 VEYAAAVPDA PPPVATAEKQ PGSNSDDELQ PEGDGDHSCS PSTSSSVHTP TSSSAL* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 212 | 217 | KRKRRR |
| 2 | 213 | 217 | RKRRR |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in phosphate signaling in roots. {ECO:0000250|UniProtKB:Q9FX67}. | |||||
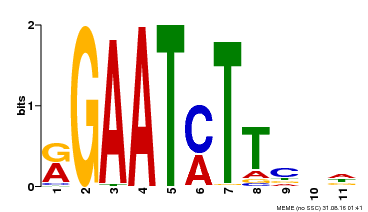
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lus10021699 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021649606.1 | 1e-122 | transcription factor HHO3-like isoform X1 | ||||
| Swissprot | Q9FPE8 | 5e-93 | HHO3_ARATH; Transcription factor HHO3 | ||||
| TrEMBL | A0A2C9VQR9 | 1e-116 | A0A2C9VQR9_MANES; Uncharacterized protein | ||||
| STRING | Lus10021699 | 0.0 | (Linum usitatissimum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3918 | 33 | 63 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25550.1 | 1e-90 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Lus10021699 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



