 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lus10015657 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Linaceae; Linum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 376aa MW: 42844.5 Da PI: 7.5836 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 19.9 | 1.9e-06 | 75 | 96 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
+C+ Cg +F+ + +L+ H+r+H
Lus10015657 75 TCEECGACFRKPAYLREHMRSH 96
6********************9 PP
| |||||||
| 2 | zf-C2H2 | 16.6 | 2.2e-05 | 102 | 126 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+kC+ C+ s++rk++L+rH+ H
Lus10015657 102 FKCTveGCPASYRRKDHLNRHLLQH 126
89*******************9887 PP
| |||||||
| 3 | zf-C2H2 | 21.9 | 4.7e-07 | 168 | 192 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
y C+ Cgk F+ +s Lk+H+ +H
Lus10015657 168 YMCEepGCGKAFPYPSKLKKHQDSH 192
899999****************998 PP
| |||||||
| 4 | zf-C2H2 | 12.7 | 0.00038 | 216 | 236 | 4 | 23 |
TTTTEEESSHHHHHHHHHH.T CS
zf-C2H2 4 pdCgksFsrksnLkrHirt.H 23
p Cgk Fs+ Lk Hi+ H
Lus10015657 216 PGCGKNFSNEQCLKAHIQAsH 236
39***************9877 PP
| |||||||
| 5 | zf-C2H2 | 16.2 | 2.9e-05 | 272 | 295 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirt.H 23
C Cg +Fs+k nL++H + H
Lus10015657 272 CDhdGCGLTFSSKTNLNKHVKAvH 295
77789**************99877 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 29 | 33 | 55 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.0048 | 74 | 96 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.796 | 74 | 101 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 3.9E-8 | 74 | 96 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 76 | 96 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 5.9E-11 | 83 | 134 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 7.8E-13 | 97 | 127 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 9.2E-4 | 102 | 126 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 8.621 | 102 | 126 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 104 | 126 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.41 | 131 | 156 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.203 | 131 | 161 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 133 | 156 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 3.13E-6 | 164 | 192 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 6.6E-9 | 164 | 194 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0011 | 168 | 192 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.344 | 168 | 192 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 170 | 192 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.15 | 211 | 236 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 5.0E-4 | 212 | 232 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 213 | 236 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 23 | 239 | 260 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 7.83E-9 | 254 | 304 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.2E-7 | 267 | 295 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 5.5E-4 | 270 | 295 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.133 | 270 | 300 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 272 | 295 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 4.4E-4 | 296 | 323 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 43 | 301 | 327 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 303 | 327 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 376 aa Download sequence Send to blast |
MEAELPEEAM EKEMEEAQDS SYTTTTTSDI RRYLCKYCGI SRSKKYLISS HILSHHKDEM 60 SDEEEEAEEG TKSNTCEECG ACFRKPAYLR EHMRSHSLER PFKCTVEGCP ASYRRKDHLN 120 RHLLQHKEKL FKCPVESCTR EFVLESDVEG HVRDKHCKSS EADGSKQYMC EEPGCGKAFP 180 YPSKLKKHQD SHGKSLSITE LLYITLDSIE AICLEPGCGK NFSNEQCLKA HIQASHQHIN 240 CEVCGSKQLK KNIKRHLRSH EAGSESTKRV SCDHDGCGLT FSSKTNLNKH VKAVHLEAKP 300 FACGFPRCGM RFSYKHVRDN HEKSGCHVYT CGDFEESDEH FRSRPRGGMK RMCPSVETLL 360 TRKRVTLPPE LENCL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2i13_A | 2e-14 | 68 | 231 | 16 | 180 | Aart |
| 2i13_B | 2e-14 | 68 | 231 | 16 | 180 | Aart |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
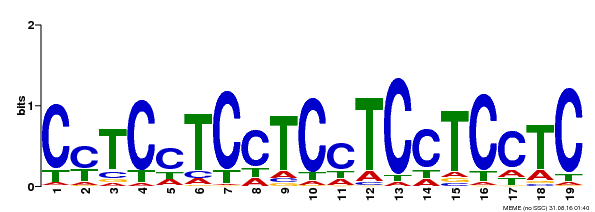
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lus10015657 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006376244.1 | 1e-163 | transcription factor IIIA | ||||
| Swissprot | Q84MZ4 | 1e-134 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | U5FVV9 | 1e-162 | U5FVV9_POPTR; Uncharacterized protein | ||||
| STRING | Lus10015657 | 0.0 | (Linum usitatissimum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3069 | 33 | 70 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-132 | transcription factor IIIA | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Lus10015657 |



