 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lus10009984 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Linaceae; Linum
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 957aa MW: 105786 Da PI: 6.508 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 93.7 | 1.8e-29 | 163 | 236 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C adls+ak+yhrrhkvCe+hska+++lv++++qrfCqqC + ++ ++ rsCrrrLa+hn+rrrk+++
Lus10009984 163 VCQVEDCGADLSKAKDYHRRHKVCEMHSKATKALVANVMQRFCQQCRK----EKLSDKMRSCRRRLAGHNKRRRKTNP 236
6*********************************************87....5667899****************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 5.3E-22 | 156 | 221 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 24.726 | 161 | 234 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.19E-27 | 162 | 238 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 1.1E-20 | 164 | 233 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 2.18E-8 | 743 | 840 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.8E-7 | 744 | 847 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 5.15E-9 | 745 | 840 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 957 aa Download sequence Send to blast |
MEARFGTEAS HFYDMRAVGK RSMEWDLNDW KWDGDLFIAR PLNAVVPSSS AAASAGMSKQ 60 FFPLNNTNNN SSNSSSSCSD EVGNLEMIEK GKRRRNKVID VEEDESLMVN GHQEEEAATL 120 SLKLGGHGGG EMVNWEGNSS GKKAKLTGGG GNNNNSSSSS RAVCQVEDCG ADLSKAKDYH 180 RRHKVCEMHS KATKALVANV MQRFCQQCRK EKLSDKMRSC RRRLAGHNKR RRKTNPEPGS 240 NSNTVNDDQT GGYLLISLLR TLSNMHSSKS EKSTDQDLLS HLLRSLAGQT IEHGNPKQAQ 300 LVHANGASLQ TSSSMKSRMP NNFPSCSEVR DSNPRSLKMN NFDLNDAYID SDDRTDDVER 360 SLVPANGGTS SLGCPSWAQQ DSLQSSPPQT SGNSDSASPQ SPSSSNGDGQ NRTDRIVFKL 420 FGKKPNDFPL VLRAQILDWL SHSPTDIESY IRPGCIVLTI YLRQDEAAWD ELCCDLSTSL 480 SRLLHTSFDS FWRTGWLFVM MEHQIAFAYN GQVVVDTSSP LRNTDCGKIC SVKPIAVSAA 540 GKVQFLIKGT NLSRSSTRIL CAIEGKYMVQ EDSDDLAHTK DIKDQGQLQC VPYSCSIPTG 600 LGRGFIEVEE DGISNSYFPF IVAEEDVCSE ICTLENALEF FETDVHGNLG EGGGKIEAKT 660 QAMEFIHEMG WLLHRIQSKS RLNDMDPNVD RFPISRMKWL MDFSMDHEWC AVVRRLLNVV 720 VRDGIVDLGG ECSSLNNALA EMGVLHRAVR RNSRPLVEFL LKYAPENDDE FLFRPDMAGP 780 GGLTPLHIAA GKDGSEDILD ALTDDPKMVG VEAWKNARDS TGFTPEDYAR LRGHYSYIHL 840 VQRKINRKLS EGSRHVVLDI PENEKQDEAL SSALEIMQSV LKKPSASSKS CKLCTGNKVA 900 YGVTSSRRSL LYRPAMLSMV AVAAVCVCVA LLFKSCPEVV YVLSPFRWEE LEYGTS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 2e-21 | 155 | 233 | 2 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
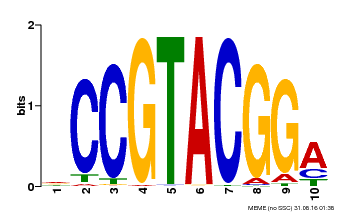
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00603 | PBM | Transfer from AT2G47070 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lus10009984 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012082979.1 | 0.0 | squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A067K935 | 0.0 | A0A067K935_JATCU; Uncharacterized protein | ||||
| STRING | Lus10009984 | 0.0 | (Linum usitatissimum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1539 | 34 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Lus10009984 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



