 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lus10006368 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Linaceae; Linum
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 507aa MW: 55387.9 Da PI: 6.2497 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 96 | 2.5e-30 | 116 | 173 | 2 | 60 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS-- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhek 60
+DgynWrKYGqK vkgsef rsYYrCt+++C+vkk++ers+ ++k+ +i+Y g+H+h+k
Lus10006368 116 EDGYNWRKYGQKLVKGSEFIRSYYRCTHPNCQVKKQLERSH-EGKLADIVYFGQHDHPK 173
8****************************************.***************85 PP
| |||||||
| 2 | WRKY | 106.6 | 1.2e-33 | 285 | 343 | 1 | 59 |
---SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 1 ldDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
++Dgy+WrKYGqK+vkg+++prsYYrC+s+gCpvkk+ver+++dpkvv++ YegeH+h+
Lus10006368 285 VNDGYRWRKYGQKMVKGNPNPRSYYRCSSPGCPVKKHVERASHDPKVVITSYEGEHDHD 343
58********************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50811 | 22.676 | 110 | 174 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 2.4E-26 | 111 | 174 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 2.35E-24 | 112 | 174 | IPR003657 | WRKY domain |
| SMART | SM00774 | 6.0E-35 | 115 | 173 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 3.1E-23 | 116 | 172 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 4.0E-36 | 271 | 345 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 3.27E-29 | 278 | 345 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 36.265 | 280 | 345 | IPR003657 | WRKY domain |
| SMART | SM00774 | 1.4E-39 | 285 | 344 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 3.0E-26 | 286 | 342 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009863 | Biological Process | salicylic acid mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 507 aa Download sequence Send to blast |
MDSSVEGPEE LPSNETQQTK SPFDSRAEKA QQIDDDKTTH TSKSQEKTMS IVVHKGAASH 60 DLDDRDCVMQ VDKEASTPSA NRGKLSLAIP TVTGGPQPLQ SVKDGRSPAI REKVLEDGYN 120 WRKYGQKLVK GSEFIRSYYR CTHPNCQVKK QLERSHEGKL ADIVYFGQHD HPKPQSNIPL 180 TVGFVLSVEE RADQCPLSPV EKGSKGRPPI SSANASQMST LTSSEDVKAV PPESARKRDE 240 IDNDDDPQSK RQKKVNRSVE PSAVVEKPVG EPRIVVKTLS EEDIVNDGYR WRKYGQKMVK 300 GNPNPRSYYR CSSPGCPVKK HVERASHDPK VVITSYEGEH DHDMPPSRTV TNNIAGLDVR 360 TSTPASGQLD SKPACNTPDA NNKTKEQLNG ESSAKAIEGT VKEAVDKGND MVSPRSSALD 420 EMKSKNQQND KSCPKEESDA AGHDSDATTS FILEGRTNDV QHTCESKETS EENGSASASN 480 IDSVASHPDS KSTQQHISDS EAVQAN* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2ayd_A | 7e-38 | 114 | 346 | 2 | 75 | WRKY transcription factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Binds to a 5'-CGTTGACCGAG-3' consensus core sequence which contains a W box, a frequently occurring elicitor-responsive cis-acting element. {ECO:0000269|PubMed:8972846}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00094 | SELEX | Transfer from AT2G04880 | Download |
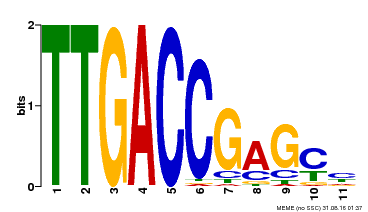
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lus10006368 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salicylic acid (SA). {ECO:0000269|PubMed:17264121}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002533869.1 | 1e-148 | WRKY transcription factor 1 | ||||
| Swissprot | Q9SI37 | 2e-87 | WRKY1_ARATH; WRKY transcription factor 1 | ||||
| TrEMBL | B9T6K0 | 1e-146 | B9T6K0_RICCO; WRKY transcription factor, putative | ||||
| STRING | Lus10006368 | 0.0 | (Linum usitatissimum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF9358 | 33 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G04880.2 | 2e-91 | zinc-dependent activator protein-1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Lus10006368 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



