 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lus10003405 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Linaceae; Linum
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1104aa MW: 123624 Da PI: 6.4517 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 165.2 | 1.1e-51 | 21 | 123 | 3 | 105 |
CG-1 3 kekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcyw 100
+ ++rwl++ ei++iL n+ek+++++e++++p sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+v+vl+cyYah+ee+++fqrr+yw
Lus10003405 21 EAQHRWLRPAEICEILRNYEKFHISTEPPNKPASGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEETESFQRRSYW 118
459*********************************************************************************************** PP
CG-1 101 lLeee 105
+Le+
Lus10003405 119 MLEHM 123
**986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 72.702 | 15 | 142 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 6.4E-66 | 18 | 137 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 2.5E-45 | 21 | 131 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 2.8E-4 | 486 | 600 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 7.0E-15 | 514 | 599 | IPR014756 | Immunoglobulin E-set |
| PROSITE profile | PS50297 | 15.777 | 698 | 809 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 6.8E-17 | 699 | 809 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 9.33E-16 | 700 | 809 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 5.85E-13 | 703 | 807 | No hit | No description |
| SuperFamily | SSF52540 | 1.12E-7 | 918 | 969 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.071 | 918 | 940 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.078 | 919 | 948 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0027 | 920 | 939 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0093 | 941 | 963 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.377 | 942 | 966 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.4E-4 | 944 | 963 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1104 aa Download sequence Send to blast |
MADRGSFGLA PRLDIQQLFI EAQHRWLRPA EICEILRNYE KFHISTEPPN KPASGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHEK LKVGSVDVLH CYYAHGEETE SFQRRSYWML 120 EHMLCFSVVL YCYFTFPSWI FLVLMALTIL QGTYAYSIRP LLGSEVLSCL YAYAGHSSLH 180 QGNRTHSSPS NSLVNHNRSG NTDSISPSST LSSAFEDADS ADSHQTTGTH SFYRPQPLQN 240 SVTIDKVDVG PSSTHLSHSR AGSGSPVAYR HGSGSGINDS SSNIINSQGL TSWEDVYEQS 300 TRGFDNTPLR AALNSTHLGN YGVGANLMQQ VSTGSTTQVN FDGSLPIHST WQIPAGDSSL 360 HIPHGMDATF DLDLTYDFGT RLFDQRTQDF GTHNTLEPYF TGSLQQHDEL LHNNLQIELL 420 NSELQYATGS NLPIEGAADG NGRYALFAKK PLLGEEGLKK VDSFSRWVSK ELGEVDDLNM 480 KSSSGIAWNN VECGNVADES VLSPSLSQDQ LFSIMDFSPK WAYADSETEV HIVGMFLKSQ 540 EEVQKCEWSC MFGEVEVRAE VLADGILCCF APPHGFARVP LYVTCSNRVA CSEVREFDYL 600 VNSVGDVNVR GVYGGNAPEM WLHLRLEKLL SVKPSTGLSH TLDVARAKQK LVRKIMLLKE 660 EDDNHQMIEP TSEESLSHVE VKRKLLYKLI KEKLYAWLLH KVLESGKGPC VLDDDGQGVL 720 HLAAALGWDW AIKPTVAAGV SINFRDVNGW TALHWAAYCG REQTVAVLVS WRAAPGALTD 780 PSPEFPSGRT AAELASGNGH KGISGFLAES FLTSYLSTLT VKEQKDDGEN VVQTIEERTA 840 TPLNGNDVPD ALSLKDSLSA IRNATQAANR IYQVFRMQSF QRKQLSEEHE SDEFGLSDEH 900 ALSLLAGNSR RPVYSNGKAH AAATQIQKKF RGWKKRREFL IIRQRIVKIQ AHIRGHQVRK 960 QYKTVVWSVG ILEKIILRWR RKGSGLRGFR RDAMIEDLPS PQNVHPKEDD DYDYLKEGRK 1020 QNEERVEKTL NRVKSMFHDP NGRAQYRRLL TVVEGLRENQ ACSSILNHPQ DETEEGQDEE 1080 SSQQLADFDS LLDDDTFMSL AFD* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
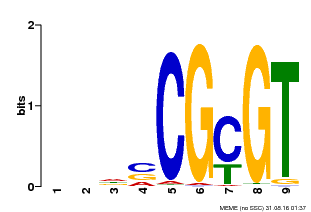
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lus10003405 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021670318.1 | 0.0 | calmodulin-binding transcription activator 2-like | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A067JX34 | 0.0 | A0A067JX34_JATCU; Uncharacterized protein | ||||
| STRING | Lus10003405 | 0.0 | (Linum usitatissimum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7406 | 26 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Lus10003405 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



