 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LPERR08G03480.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Leersia
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 804aa MW: 87974.8 Da PI: 7.0626 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 47.7 | 3.7e-15 | 107 | 151 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+ E++++++a k++G W +I +++g ++t+ q++s+ qk+
LPERR08G03480.1 107 RERWTEAEHKRFLEALKLYGRA-WQRIEEHVG-TKTAVQIRSHAQKF 151
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.29E-16 | 101 | 157 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.147 | 102 | 156 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.9E-16 | 105 | 154 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 6.5E-12 | 106 | 154 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.7E-12 | 107 | 150 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.0E-8 | 107 | 147 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.20E-8 | 109 | 152 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010243 | Biological Process | response to organonitrogen compound | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043496 | Biological Process | regulation of protein homodimerization activity | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 804 aa Download sequence Send to blast |
MTMLLSRRRR GRPWPACGGL GLLRGASQRA RRQRTRFDST GLLCLLEVCD HASRATAVGW 60 HLFVVLCCVR LKEVSGLSRL TAIGSIQINS SGEEAMVRKP YTITKQRERW TEAEHKRFLE 120 ALKLYGRAWQ RIEEHVGTKT AVQIRSHAQK FFTKLEKEAI NNGTSPGQAH DIDIPPPRPK 180 RKPNSPYPRK SCLSSETSTR EIPNDKATKS NLTVNSTAQM AGDAALEKLQ RKEISEKGSC 240 SEVLNLFREA PSASFSSVNK SSSNHGASRG LEPTKTEIKD VVILERDGIS NGAGKNVKDI 300 NDQETERLNG IHISSKSDHS HENCLDTSRQ QLKPKATAVE PTYVDWSAAK TSHCQMDRNG 360 ATGFPATGTE GSHPDQTSDQ TGRTSETMNQ CIHPTLSVDP KFDSTATAQP FPHNYAAFAP 420 MMQCHCNQDS YRSFVNMSST FSSMLVSTLL SNPAIHAAAR LAASYWPAVD GNSADPNQED 480 LSESAQGSHA GSPPNMASIV AATVAAASAW WATQGLLPLF PPPIAFPFVP APSTPFPAAD 540 VQRAPEKDMN CPIDSAQKEM QETQKQDNPE AIKVIMSSES DESGKGEVSL HTELKISPAD 600 KADATPTTGA DTSDVFGNKK KQDRSSCGSN TPSSSDHDIE ADNAPEKQEK ANDKAKQASC 660 SNSSAGDNNH RRFRSSGSTS DSWKEVSEEG RLAFDALFSR ERLPQSFSPP QAEESKEISK 720 EEEDEVTTVT VDLNKNATII DHELDTVDDP RAFFPNELSN LKLKSRRTGF KPYKRCSVEA 780 KENRVPASDE VGTKRIRLDS ETST |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 28 | 35 | RARRQRTR |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
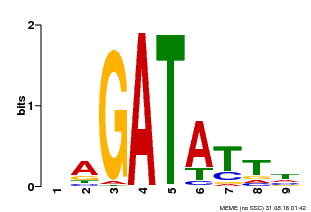
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LPERR08G03480.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK242771 | 0.0 | AK242771.1 Oryza sativa Japonica Group cDNA, clone: J090053J23, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015649844.1 | 0.0 | protein LHY isoform X2 | ||||
| Refseq | XP_015649845.1 | 0.0 | protein LHY isoform X2 | ||||
| TrEMBL | A0A0D9X4L5 | 0.0 | A0A0D9X4L5_9ORYZ; Uncharacterized protein | ||||
| STRING | LPERR08G03480.1 | 0.0 | (Leersia perrieri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6031 | 34 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01060.4 | 4e-37 | MYB_related family protein | ||||



