 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LPERR06G21800.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Leersia
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 358aa MW: 38387.2 Da PI: 4.5395 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 51.3 | 1.6e-16 | 275 | 311 | 1 | 35 |
GATA 1 CsnCgttk..TplWRrgpdgnktLCnaCGlyyrkkgl 35
C++Cg + Tp++Rrgpdg++tLCnaCGl+++ kg+
LPERR06G21800.1 275 CHHCGISAkaTPMMRRGPDGPRTLCNACGLMWANKGM 311
*****88889***********************9996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51320 | 15.336 | 139 | 174 | IPR010399 | Tify domain |
| SMART | SM00979 | 5.0E-12 | 139 | 174 | IPR010399 | Tify domain |
| Pfam | PF06200 | 4.1E-13 | 142 | 173 | IPR010399 | Tify domain |
| PROSITE profile | PS51017 | 13.404 | 200 | 242 | IPR010402 | CCT domain |
| Pfam | PF06203 | 8.4E-16 | 200 | 242 | IPR010402 | CCT domain |
| SMART | SM00401 | 1.3E-11 | 269 | 322 | IPR000679 | Zinc finger, GATA-type |
| PROSITE profile | PS50114 | 9.222 | 269 | 326 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 2.33E-12 | 273 | 325 | No hit | No description |
| CDD | cd00202 | 3.22E-14 | 274 | 325 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 5.3E-16 | 274 | 316 | IPR013088 | Zinc finger, NHR/GATA-type |
| PROSITE pattern | PS00344 | 0 | 275 | 302 | IPR000679 | Zinc finger, GATA-type |
| Pfam | PF00320 | 4.3E-14 | 275 | 310 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 358 aa Download sequence Send to blast |
MSHHDGSKPY QPRRRPERPP PPPPPPDDAA HSGPSVDDLA AAAAASEAEA IKRYQERYGM 60 EVAEGEEDLE EEELEEEEEE EEMEEEEMGD EEEQQHHEGG GAEEEAVPMD AEAAAAAVAA 120 MGGQMDQHGT MVAAAAAVPP MASNQLTLSF QGEVYVFDSV SPDKVQAVLL LLGGRELNPG 180 LGSGASSSTS YSKRLNFPHR VASLMRFREK RKERNFDKKI RYSVRKEVAL RMQRNRGQFT 240 SSKTKGDEAT SELTASDGSP NWGSVEGRPP SAAECHHCGI SAKATPMMRR GPDGPRTLCN 300 ACGLMWANKG MLRDLSKAPP TPIQAVPSAN DGNGSDAAPA TEQEIAAPAT ANGHDSST |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds 5'-GATA-3' or 5'-GAT-3' motifs within gene promoters. {ECO:0000250|UniProtKB:Q8LAU9}. | |||||
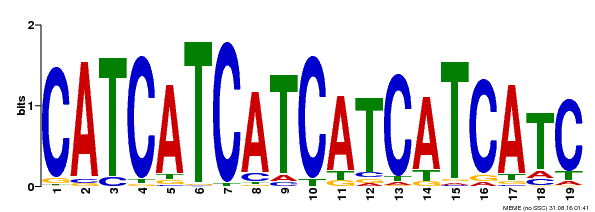
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00198 | DAP | Transfer from AT1G51600 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LPERR06G21800.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By abscisic acid (ABA), and drought and salt stresses. Down-regulated by jasmonate and wounding. {ECO:0000269|PubMed:19618278}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK243334 | 0.0 | AK243334.1 Oryza sativa Japonica Group cDNA, clone: J100058P16, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025882282.1 | 1e-173 | GATA transcription factor 20 | ||||
| Swissprot | Q5Z4U5 | 1e-175 | GAT20_ORYSJ; GATA transcription factor 20 | ||||
| TrEMBL | A0A0D9WTQ4 | 0.0 | A0A0D9WTQ4_9ORYZ; Uncharacterized protein | ||||
| STRING | LPERR06G21800.1 | 0.0 | (Leersia perrieri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4325 | 38 | 68 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G51600.2 | 2e-75 | ZIM-LIKE 2 | ||||



