 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LPERR06G01660.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Leersia
|
||||||||
| Family | BBR-BPC | ||||||||
| Protein Properties | Length: 331aa MW: 36685.4 Da PI: 9.9794 | ||||||||
| Description | BBR-BPC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GAGA_bind | 402.8 | 4.5e-123 | 1 | 331 | 1 | 301 |
GAGA_bind 1 mdddgsre..rnkg.yye..........paaslkenlglqlmssiaerdakirernlalsekkaavaerdmaflqrdkalaernkalverdnkl 81
md+ g+re r+++ +y+ +++lk++++++l++++++rd++irer++al+ekkaa+aerdmaf+qrd+a+aern+a+verdn+l
LPERR06G01660.1 1 MDNLGHREngRQRPdQYKglhtqwmmpqTQRHLKDHQSMNLLALMNDRDNAIRERDHALAEKKAAIAERDMAFAQRDAAMAERNAAVVERDNAL 94
9999999999****9***********74449*************************************************************** PP
GAGA_bind 82 lalllvensla...salpvgvqvlsgtksidslqqlse.....pqledsave.lreeeklealpieeaaeeakekkkkkkrqrakkpkekkak. 165
+al+l++++ + ++ +++l+g+k+i++++qls+ ql+ds+++ +re++++ea+pi++a+ ++ ++k++kk++++++p+++ +
LPERR06G01660.1 95 AALELARTNGLnmnNGNGYPQGSLNGSKNIHHHDQLSHaqaspLQLADSPYDhAREMHISEAYPISTAPGSVGKAKRPKKNSSQASPLKRPSGv 188
***9988876555577888999*************888899999********9*************************************9998 PP
GAGA_bind 166 .kkkkksekskkkvkkesader......skaekksidlvlngvslDestlPvPvCsCtGalrqCYkWGnGGWqSaCCtttiSvyPLPvstkrrg 252
+k+kk+++++k+v ++ + ++ +k+++k+ dl+ln+v++Dest+P+P+CsCtG+lrqCYkWGnGGWqS+CCt++iS+yPLPv +++r+
LPERR06G01660.1 189 lRKTKKPSSDWKNVGMSDCGDDsanasvMKNDWKDKDLGLNQVAFDESTMPAPACSCTGKLRQCYKWGNGGWQSSCCTMNISMYPLPVIPNKRH 282
88888899********999854589999****************************************************************** PP
GAGA_bind 253 aRiagrKmSqgafkklLekLaaeGydlsnpvDLkdhWAkHGtnkfvtir 301
aR++grKmS+gaf+klL++LaaeG+dls+pvDLkdhWAkHGtn+++tir
LPERR06G01660.1 283 ARMGGRKMSGGAFTKLLSRLAAEGHDLSTPVDLKDHWAKHGTNRYITIR 331
************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01226 | 3.3E-180 | 1 | 331 | IPR010409 | GAGA-binding transcriptional activator |
| Pfam | PF06217 | 3.1E-97 | 1 | 331 | IPR010409 | GAGA-binding transcriptional activator |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 331 aa Download sequence Send to blast |
MDNLGHRENG RQRPDQYKGL HTQWMMPQTQ RHLKDHQSMN LLALMNDRDN AIRERDHALA 60 EKKAAIAERD MAFAQRDAAM AERNAAVVER DNALAALELA RTNGLNMNNG NGYPQGSLNG 120 SKNIHHHDQL SHAQASPLQL ADSPYDHARE MHISEAYPIS TAPGSVGKAK RPKKNSSQAS 180 PLKRPSGVLR KTKKPSSDWK NVGMSDCGDD SANASVMKND WKDKDLGLNQ VAFDESTMPA 240 PACSCTGKLR QCYKWGNGGW QSSCCTMNIS MYPLPVIPNK RHARMGGRKM SGGAFTKLLS 300 RLAAEGHDLS TPVDLKDHWA KHGTNRYITI R |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds to GA-rich elements (GAGA-repeats) present in regulatory sequences of genes involved in developmental processes. {ECO:0000250}. | |||||
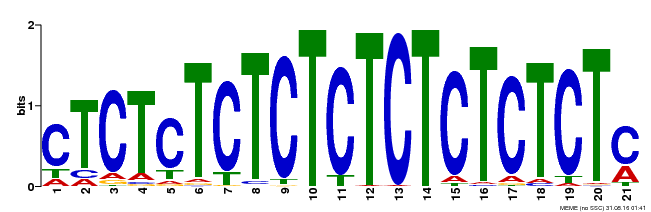
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00540 | DAP | Transfer from AT5G42520 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LPERR06G01660.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK070616 | 0.0 | AK070616.1 Oryza sativa Japonica Group cDNA clone:J023060O20, full insert sequence. | |||
| GenBank | AY957505 | 0.0 | AY957505.1 Oryza sativa (japonica cultivar-group) barley B recombinant-like protein D mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015641153.1 | 0.0 | barley B recombinant-like protein D | ||||
| Refseq | XP_015641154.1 | 0.0 | barley B recombinant-like protein D | ||||
| Swissprot | Q5VSA8 | 0.0 | BBRD_ORYSJ; Barley B recombinant-like protein D | ||||
| TrEMBL | A0A0D9WLD5 | 0.0 | A0A0D9WLD5_9ORYZ; Uncharacterized protein | ||||
| STRING | LPERR06G01660.1 | 0.0 | (Leersia perrieri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6226 | 37 | 56 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42520.1 | 1e-102 | basic pentacysteine 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



