 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LPERR05G14660.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Leersia
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 307aa MW: 33672.5 Da PI: 6.5738 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 29.5 | 1.7e-09 | 31 | 76 | 1 | 44 |
TSSS-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrw 44
rg+WT+eE + + +a + + +W+++a++++ g+t ++ ++
LPERR05G14660.1 31 RGAWTPEENKAFEQALAAVDRNdpeRWERVAEMLP-GKTVADVMTHY 76
89*************9999988999**********.********999 PP
| |||||||
| 2 | Myb_DNA-binding | 45.7 | 1.5e-14 | 148 | 192 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE++l++ + k++G g+W+ I+r + + Rt+ q+ s+ qky
LPERR05G14660.1 148 PWTEEEHKLFLMGLKKYGRGDWRNISRNFVTSRTPTQVASHAQKY 192
8*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-5 | 30 | 78 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50090 | 7.039 | 30 | 80 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 2.1E-9 | 30 | 82 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.1E-7 | 31 | 76 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 1.72E-10 | 32 | 87 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.40E-7 | 33 | 76 | No hit | No description |
| PROSITE profile | PS51294 | 17.568 | 141 | 197 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.36E-17 | 143 | 197 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.8E-17 | 144 | 195 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 2.0E-12 | 145 | 191 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 5.8E-13 | 145 | 195 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 7.1E-12 | 148 | 192 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.95E-11 | 148 | 193 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 307 aa Download sequence Send to blast |
MESYMEVLPP APAHYFVGQP SAGGWFLQDR RGAWTPEENK AFEQALAAVD RNDPERWERV 60 AEMLPGKTVA DVMTHYDDLE NDVCFIEAGL VPFPQYGAGA GSPSSGFTFD WDGADDAAGL 120 AFKRSCYIVG GGKRGARGPD QERKKGVPWT EEEHKLFLMG LKKYGRGDWR NISRNFVTSR 180 TPTQVASHAQ KYFIRLNSGG KDKRRSSIHD ITTVNLPDED HGNNPSPSPP SVLTAAHSSS 240 SSAAVSDQFG VLVDSKPPPP TTLGHHHFMP HHPYAQVKLE ASNSHVGGRF DDSVLMQMQC 300 GQLQPLG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 5e-15 | 34 | 102 | 11 | 79 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the dorsovental asymmetry of flowers. Promotes ventral identity. {ECO:0000269|PubMed:11937495, ECO:0000269|PubMed:9118809}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00296 | DAP | Transfer from AT2G38090 | Download |
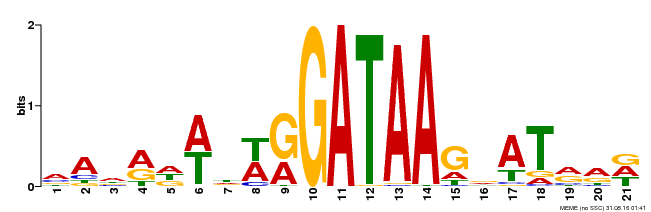
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LPERR05G14660.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK068138 | 0.0 | AK068138.1 Oryza sativa Japonica Group cDNA clone:J013135D01, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006654489.1 | 0.0 | PREDICTED: transcription factor DIVARICATA-like | ||||
| Swissprot | Q8S9H7 | 6e-82 | DIV_ANTMA; Transcription factor DIVARICATA | ||||
| TrEMBL | A0A0D9WH62 | 0.0 | A0A0D9WH62_9ORYZ; Uncharacterized protein | ||||
| STRING | LPERR05G14660.1 | 0.0 | (Leersia perrieri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2274 | 37 | 96 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38090.1 | 2e-78 | MYB family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



