 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LPERR03G08040.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Leersia
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 685aa MW: 73401.5 Da PI: 6.4671 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 89.9 | 2.2e-28 | 202 | 255 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv+av++L G++kA+Pk+ilelm+v+gLt+e+v+SHLQk+Rl
LPERR03G08040.2 202 KPRVVWSVELHQQFVNAVNHL-GIDKAVPKKILELMNVPGLTRENVASHLQKFRL 255
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 78.4 | 2.5e-26 | 18 | 126 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHT CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealka 92
vl+vdD+p+ + +l+++l + +y +++++++++ al++l+e++ +D+i+ D+ mp+mdG++ll+ e++lp+i+++a + + + + +k
LPERR03G08040.2 18 VLVVDDDPTCLAVLKRMLLECRY-DATTCSQATRALTMLRENRrgYDVIISDVHMPDMDGFRLLELVGL-EMDLPVIMMSADSRTDIVMKGIKH 109
89*********************.***************99999********************87754.558********************* PP
TESEEEESS--HHHHHH CS
Response_reg 93 GakdflsKpfdpeelvk 109
Ga d+l Kp+ +eel +
LPERR03G08040.2 110 GACDYLIKPVRMEELKN 126
**************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 1.3E-192 | 1 | 668 | IPR017053 | Response regulator B-type, plant |
| Gene3D | G3DSA:3.40.50.2300 | 4.3E-42 | 15 | 150 | No hit | No description |
| SuperFamily | SSF52172 | 7.54E-36 | 15 | 142 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 5.5E-30 | 16 | 128 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 42.948 | 17 | 132 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 5.4E-23 | 18 | 126 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 3.65E-27 | 19 | 131 | No hit | No description |
| SuperFamily | SSF46689 | 4.57E-20 | 199 | 259 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 11.674 | 199 | 258 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-29 | 199 | 260 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.0E-25 | 202 | 255 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 7.8E-7 | 204 | 253 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0031537 | Biological Process | regulation of anthocyanin metabolic process | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0080036 | Biological Process | regulation of cytokinin-activated signaling pathway | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 685 aa Download sequence Send to blast |
MAPVDDGGGG EFPVGMKVLV VDDDPTCLAV LKRMLLECRY DATTCSQATR ALTMLRENRR 60 GYDVIISDVH MPDMDGFRLL ELVGLEMDLP VIMMSADSRT DIVMKGIKHG ACDYLIKPVR 120 MEELKNIWQH VVRKKFNESK EHEHSGSLDD TDRTRPANND NEYASSANDG ADGSWKSQKK 180 KRDKDEDDGE LESGDPSSTS KKPRVVWSVE LHQQFVNAVN HLGIDKAVPK KILELMNVPG 240 LTRENVASHL QKFRLYLKRI AQHHAGIANP FCPPASNAKV GSLGGLDFQA LAASGQIPPQ 300 ALAALQDELL GRPTNSLVLP GRDQSSLRLA AVKGNKPHGE REIAFGQPIY KCQNNAYGAF 360 PQSSPTVGGM PSFSAWPNNK LGMADSTGTL GSMSNSQNSN IVLHELQQQP DAMLSGTIHS 420 LDVKPSGIVP SQSLNTFSTS EGLSPNQNTL MIPTQSSGFL AAMPPSMKHE PVLATSQPSS 480 SLLGGIDLVN QASTSQSLMS THGGLVNRNP NVIPSQGIST FQTTNNPYLV NPNSMGVGSK 540 QQTGVLKTEN SDTLSHSYGY LGGNTPPMDS CLLSSQSKNT QFGLLGQDDI TGSWSPLPNI 600 DSFGNTVGLS HPGSGSSSFQ SSNVALGKLP DQGRGKNHGF VGKGTCIPSR FAVDEIESPP 660 NNLSHSIGSS GDIMSPDIFG FSGQM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 7e-24 | 198 | 261 | 1 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specific DNA sequence. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. May directly activate some type-A response regulators in response to cytokinins. {ECO:0000250|UniProtKB:Q940D0}. | |||||
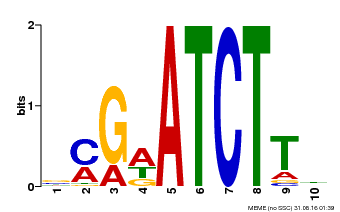
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LPERR03G08040.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB246780 | 0.0 | AB246780.1 Oryza sativa Japonica Group ORR1 mRNA for B-type response regulator, complete cds. | |||
| GenBank | AK111864 | 0.0 | AK111864.1 Oryza sativa Japonica Group cDNA clone:J033025G23, full insert sequence. | |||
| GenBank | AK111899 | 0.0 | AK111899.1 Oryza sativa Japonica Group cDNA clone:J023034P21, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015630577.1 | 0.0 | two-component response regulator ORR21 | ||||
| Refseq | XP_015630578.1 | 0.0 | two-component response regulator ORR21 | ||||
| Swissprot | A2XE31 | 0.0 | ORR21_ORYSI; Two-component response regulator ORR21 | ||||
| TrEMBL | A0A0D9VRE8 | 0.0 | A0A0D9VRE8_9ORYZ; Two-component response regulator | ||||
| STRING | LPERR03G08040.1 | 0.0 | (Leersia perrieri) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G16857.1 | 1e-117 | response regulator 1 | ||||



