 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LPERR02G22520.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Leersia
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 802aa MW: 86082.3 Da PI: 5.6138 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 66.6 | 3.3e-21 | 97 | 152 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r+eL+++l+L+ rqVk+WFqNrR+++k
LPERR02G22520.1 97 KKRYHRHTPQQIQELEALFKECPHPDEKQRAELSRRLSLDARQVKFWFQNRRTQMK 152
678899***********************************************999 PP
| |||||||
| 2 | START | 188.6 | 3.2e-59 | 312 | 544 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE......EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHH.HHHHHHCCCGGCT-TT-S....E CS
START 1 elaeeaaqelvkkalaeepgWvkss......esengdevlqkfeeskv.....dsgealrasgvvdmvla.llveellddkeqWdetla....k 78
ela +a++elvk+a+ e+p+Wv e +n +e+l +f + + + +ea+r+sg+v+ + lve+l+d + +W+ ++ k
LPERR02G22520.1 312 ELAISAMDELVKMAQMEDPLWVPALpgapskEVLNFEEYLHSFIPCIGmkpagYVSEASRESGLVIIDNSlALVETLMDER-RWSDMFScmiaK 404
57889*****************99988888888888888888877777*****************9876658999999999.************ PP
EEEEEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE..-TTS--....-TTSEE-EESSEEEEEEEECTC CS
START 79 aetlevissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvd..seqkppe...sssvvRaellpSgiliepksng 160
a++le +s+g gal lm+aelq+lsplvp R+++f+R+++ql +g w++vdvS+d ++++++ +++ v++++lpSg+++++++ng
LPERR02G22520.1 405 ATVLEEVSTGiagsrnGALLLMKAELQVLSPLVPiREVTFLRFCKQLAEGAWAVVDVSIDglVRDHNSAtapTAGNVKCRRLPSGCVMQDTPNG 498
***********************************************************96545666657789********************* PP
EEEEEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 161 hskvtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
++kvtwveh+++++ ++h+l+r+l++sgla+ga++w+atlqrqce+
LPERR02G22520.1 499 YCKVTWVEHTEYDEVSVHQLYRPLLRSGLAFGARRWLATLQRQCEC 544
********************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 4.7E-22 | 81 | 148 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 6.68E-21 | 87 | 154 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 18.123 | 94 | 154 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.5E-19 | 95 | 158 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 5.33E-19 | 96 | 154 | No hit | No description |
| Pfam | PF00046 | 9.5E-19 | 97 | 152 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 129 | 152 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 45.186 | 303 | 547 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 6.11E-32 | 306 | 544 | No hit | No description |
| CDD | cd08875 | 5.13E-116 | 307 | 543 | No hit | No description |
| SMART | SM00234 | 1.7E-47 | 312 | 544 | IPR002913 | START domain |
| Pfam | PF01852 | 1.7E-51 | 313 | 544 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 6.11E-21 | 565 | 768 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048497 | Biological Process | maintenance of floral organ identity | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 802 aa Download sequence Send to blast |
MSFGGLFDGG GGGGMQFPFT SGFSSSPALS LALDNAGGGI GGRMLAGGAG SSAGGAVTRD 60 TEAENDSRSG SDHLDAISAA GEDDVEDAEP SNSRKRKKRY HRHTPQQIQE LEALFKECPH 120 PDEKQRAELS RRLSLDARQV KFWFQNRRTQ MKTQLERHEN ALLKQENDKL RAENMTIREA 180 MRSPMCGSCG SPAMLGEVSL EEQHLRIENA RLKDELNRVC ALATKFLGKP VNLMSPPPFP 240 QPHLSFPMPN SSLELAIGGI GGLGSLGTLP GCMGEFAGGV SSPMGTVITP ARATGPALPS 300 LVGNIDRSVF LELAISAMDE LVKMAQMEDP LWVPALPGAP SKEVLNFEEY LHSFIPCIGM 360 KPAGYVSEAS RESGLVIIDN SLALVETLMD ERRWSDMFSC MIAKATVLEE VSTGIAGSRN 420 GALLLMKAEL QVLSPLVPIR EVTFLRFCKQ LAEGAWAVVD VSIDGLVRDH NSATAPTAGN 480 VKCRRLPSGC VMQDTPNGYC KVTWVEHTEY DEVSVHQLYR PLLRSGLAFG ARRWLATLQR 540 QCECLAILMS STTVAANDST AISQEGKRSM LKLARRMTEN FCAGVSASSA REWSKLDGAT 600 GSIGEDVRVM ARKSVSEPGE PPGVVLSAAT SVWVPVAPEK LFNFLRDEQL RAEWDILSNG 660 GPMQEMTQIA KGQRDGNSVS LLRASAVSAN QSSMLILQET CTDASGSIVV YAPVDIPAMQ 720 LVMNGGDSTY VALLPSGFAI LPNGPRIGTS GYETGGSLLT VAFQILVNNQ PTAKLTVESV 780 ETVNNLISCT IKKIKTALQC DA |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 93 | 97 | RKRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
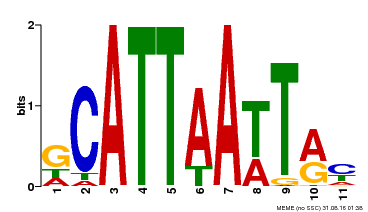
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LPERR02G22520.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB101648 | 0.0 | AB101648.1 Oryza sativa Japonica Group Roc5 mRNA for GL2-type homeodomain protein, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006647677.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ROC5 | ||||
| Swissprot | Q6EPF0 | 0.0 | ROC5_ORYSJ; Homeobox-leucine zipper protein ROC5 | ||||
| TrEMBL | A0A0D9VJF6 | 0.0 | A0A0D9VJF6_9ORYZ; Uncharacterized protein | ||||
| STRING | LPERR02G22520.1 | 0.0 | (Leersia perrieri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1203 | 37 | 116 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



