 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LPERR01G37140.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Leersia
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 804aa MW: 90494.2 Da PI: 6.5914 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 74 | 1.7e-23 | 130 | 231 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr 87
f+k+lt sd++++g +++ +++a+e+ + ++ +++l+ +d++g+ W++++i+r++++r++l++GW+ Fv++++L +gD ++F +
LPERR01G37140.1 130 FCKTLTASDTSTHGGFSVLRRHADEClppldmsQ--SPPTQELVAKDLHGNDWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL--RG 219
99*****************************644..444559************************************************..44 PP
SEE..EEEEE-S CS
B3 88 sefelvvkvfrk 99
+++el+v+v+r+
LPERR01G37140.1 220 ENGELRVGVRRA 231
9999*****996 PP
| |||||||
| 2 | Auxin_resp | 110.8 | 1.4e-36 | 256 | 337 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha++tks+F+v+Y+Pr+s+seF+++++++++++k+++svGmRf+m+fe+e+++e+r++Gt++g ++ld+ Wp+S WrsLk
LPERR01G37140.1 256 AWHAINTKSMFTVYYKPRTSPSEFIIPYDQYMESVKNNYSVGMRFRMRFEGEEAPEQRFTGTIIGSENLDQ-LWPESSWRSLK 337
89******************************************************************999.9********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 1.57E-40 | 122 | 259 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 5.5E-41 | 123 | 245 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 5.87E-21 | 128 | 230 | No hit | No description |
| SMART | SM01019 | 3.8E-22 | 130 | 232 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.899 | 130 | 232 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.9E-21 | 130 | 231 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.8E-33 | 256 | 337 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 1.0E-14 | 674 | 779 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 3.38E-13 | 690 | 771 | No hit | No description |
| PROSITE profile | PS51745 | 26.502 | 694 | 787 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 804 aa Download sequence Send to blast |
MPPAAMAPPP PPQHGSSTGD PLYDELWHAC AGPLVTVPRI GDLVFYFPQG HIEQVEASMN 60 QVADNQMRLY DLPSKLLCRV LNVELKAEQD TDEVYAQVML MPEPEQNEMA VEKSTPTSGP 120 VQARPPVRSF CKTLTASDTS THGGFSVLRR HADECLPPLD MSQSPPTQEL VAKDLHGNDW 180 RFRHIFRGQP RRHLLQSGWS VFVSSKRLVA GDAFIFLRGE NGELRVGVRR AMRQMSNVPS 240 SVISSQSMHL GVLATAWHAI NTKSMFTVYY KPRTSPSEFI IPYDQYMESV KNNYSVGMRF 300 RMRFEGEEAP EQRFTGTIIG SENLDQLWPE SSWRSLKVRW DEPSNIPRPD RVSPWKIEPA 360 SSPPVNPLPL SRVKRPRPNA PPASPESPVL TKEAATKVDI DSAQVRSQNS MVLQGQEQMT 420 LRNSLTDSND SDATVHKPMI WSPSPNGAKA HPLTFQQRPS IPMDSWMQLG RRETDFKDAR 480 SVSQSFGDSP GFFMQNFDEP SNHHASFKNQ FQDQGSARHF SDPYLYVPSQ HSLTVESSTQ 540 MHTDSKELHF WNGQSTVYGN SRDQPQNFRF EQNSSSWLNQ PFTRPEQPRV IRPHASIAPV 600 ELEKTEGSGF KIFGFKVDTT NTPNNHLSSP MAATHEPMLQ TPASLNQLQP AQTDCVPEVS 660 VSTAGTTTEN EKSAQQAQQS SKDVQNKSQG ASTRSCTKVH KQGVALGRSV DLSKFNNYDE 720 LKAELDKMFE FEGELVSSNK NWQIVYTDNE GDMMLVGDDP WEEFCSIVRK IYIYTKEEVQ 780 KMNSKSNTPR KEDSSENEKG SVKR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-170 | 15 | 362 | 12 | 357 | Auxin response factor 1 |
| 4ldw_A | 1e-170 | 15 | 362 | 12 | 357 | Auxin response factor 1 |
| 4ldw_B | 1e-170 | 15 | 362 | 12 | 357 | Auxin response factor 1 |
| 4ldx_A | 1e-170 | 15 | 362 | 12 | 357 | Auxin response factor 1 |
| 4ldx_B | 1e-170 | 15 | 362 | 12 | 357 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 373 | 377 | KRPRP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
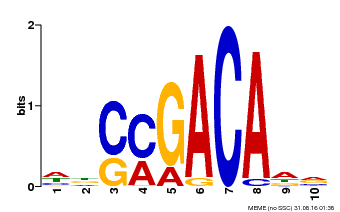
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LPERR01G37140.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK072309 | 0.0 | AK072309.1 Oryza sativa Japonica Group cDNA clone:J023022C23, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015650953.1 | 0.0 | auxin response factor 4 | ||||
| Swissprot | Q5JK20 | 0.0 | ARFD_ORYSJ; Auxin response factor 4 | ||||
| TrEMBL | A0A0D9V9T5 | 0.0 | A0A0D9V9T5_9ORYZ; Auxin response factor | ||||
| STRING | LPERR01G37140.1 | 0.0 | (Leersia perrieri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1178 | 36 | 111 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



