 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LPERR01G23470.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Leersia
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 381aa MW: 40248.4 Da PI: 9.6443 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 36.8 | 9.6e-12 | 73 | 121 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
s++kGV + +grW A+I++ r +r++lg+f + +Aa+a++ a+++++g
LPERR01G23470.1 73 SKFKGVVPQP-NGRWGAQIYE------RhQRVWLGTFAGEADAARAYDVAAQRFRG 121
79****9888.8*********......44**********99*************98 PP
| |||||||
| 2 | B3 | 98.4 | 4.3e-31 | 207 | 302 | 1 | 87 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-.. CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkld.. 85
f+k+ tpsdv+k++rlv+pk++ae+h g + +++ l +ed g++W+++++y+++s++yvltkGW++Fvk++gL +gD+v F++
LPERR01G23470.1 207 FDKTVTPSDVGKLNRLVIPKQHAEKHfplqlpsAGGESKGVLLNFEDAAGKVWRFRYSYWNSSQSYVLTKGWSRFVKEKGLHAGDVVGFYRAaa 300
89*******************************5556799*************************************************94422 PP
SS CS
B3 86 gr 87
g+
LPERR01G23470.1 301 GD 302
44 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 4.84E-22 | 73 | 129 | No hit | No description |
| SuperFamily | SSF54171 | 7.85E-16 | 73 | 129 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 1.6E-7 | 73 | 121 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 9.2E-25 | 74 | 135 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.203 | 74 | 129 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 6.7E-19 | 74 | 129 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 2.2E-38 | 201 | 312 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.72E-29 | 205 | 310 | No hit | No description |
| SuperFamily | SSF101936 | 2.35E-30 | 205 | 299 | IPR015300 | DNA-binding pseudobarrel domain |
| SMART | SM01019 | 1.1E-21 | 207 | 306 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.578 | 207 | 316 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 5.6E-28 | 207 | 307 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 381 aa Download sequence Send to blast |
MDSSSSLVDD TNSGGGGSGS SSTDKLRALV AAAAETAPLE RMGSGASAVV DAAEPGAEAD 60 SGGRVVGGKL PSSKFKGVVP QPNGRWGAQI YERHQRVWLG TFAGEADAAR AYDVAAQRFR 120 GRDAVTNFRP LADADPDAAA ELRFLAARSK AEVVDMLRKH TYFDELAQSK RALLAAFTPS 180 SAATTTPAAS LIPSPRSPAA AARQHLFDKT VTPSDVGKLN RLVIPKQHAE KHFPLQLPSA 240 GGESKGVLLN FEDAAGKVWR FRYSYWNSSQ SYVLTKGWSR FVKEKGLHAG DVVGFYRAAA 300 GDDDGGKLFI DCKLVRPTSA AVADQAAASP AVKNSIRLFG VDLLMAPTPD EHLAGCKRAR 360 DLATTPQAAA LKKQCIELAL V |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 4e-48 | 206 | 313 | 13 | 115 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00025 | PBM | Transfer from AT1G68840 | Download |
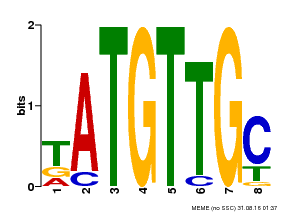
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LPERR01G23470.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK241984 | 0.0 | AK241984.1 Oryza sativa Japonica Group cDNA, clone: J075097P17, full insert sequence. | |||
| GenBank | AP003450 | 0.0 | AP003450.3 Oryza sativa Japonica Group genomic DNA, chromosome 1, PAC clone:P0034C09. | |||
| GenBank | AP014957 | 0.0 | AP014957.1 Oryza sativa Japonica Group DNA, chromosome 1, cultivar: Nipponbare, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015622542.1 | 0.0 | AP2/ERF and B3 domain-containing protein Os01g0693400 | ||||
| Swissprot | Q8RZX9 | 0.0 | Y1934_ORYSJ; AP2/ERF and B3 domain-containing protein Os01g0693400 | ||||
| TrEMBL | A0A0D9V4F9 | 0.0 | A0A0D9V4F9_9ORYZ; Uncharacterized protein | ||||
| STRING | LPERR01G23470.1 | 0.0 | (Leersia perrieri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP456 | 38 | 203 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G68840.2 | 1e-100 | related to ABI3/VP1 2 | ||||



