 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lj6g3v2218220.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Loteae; Lotus
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1041aa MW: 117088 Da PI: 5.6045 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 180.9 | 1.5e-56 | 18 | 134 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfq 95
+e ++rwl++ ei++iL n++ +++t+e++++p+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+v+vl+cyYah+een++fq
Lj6g3v2218220.1 18 REaQNRWLRPAEICEILRNYRMFDITSEPHNKPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEENENFQ 111
45599***************************************************************************************** PP
CG-1 96 rrcywlLeeelekivlvhylevk 118
rr+yw+Le+++ +iv+vhylevk
Lj6g3v2218220.1 112 RRSYWMLEQDMMHIVFVHYLEVK 134
********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 79.837 | 13 | 139 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 4.4E-79 | 16 | 134 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 2.1E-50 | 19 | 133 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 8.9E-6 | 455 | 543 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 4.7E-5 | 457 | 536 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 1.3E-16 | 457 | 543 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:1.25.40.20 | 3.7E-21 | 624 | 750 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 4.0E-8 | 634 | 714 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 20.765 | 638 | 759 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.87E-20 | 644 | 749 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.59E-16 | 644 | 747 | No hit | No description |
| SMART | SM00248 | 1300 | 655 | 684 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 9.484 | 655 | 687 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 10.873 | 688 | 720 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0015 | 688 | 717 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 1900 | 727 | 756 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 0.27 | 863 | 885 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.187 | 864 | 893 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 1.83E-7 | 865 | 914 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| Pfam | PF00612 | 0.0031 | 866 | 884 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0071 | 886 | 908 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.413 | 887 | 911 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.2E-4 | 889 | 908 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1041 aa Download sequence Send to blast |
MSEPASYGPR LDLQELQREA QNRWLRPAEI CEILRNYRMF DITSEPHNKP PSGSLFLFDR 60 KVLRYFRKDG HNWRKKKDGK TVKEAHEKLK VGSVDVLHCY YAHGEENENF QRRSYWMLEQ 120 DMMHIVFVHY LEVKGNKSNI GGNTNSDGVI SDSQVNSPSS GFPASYSSVP SLSTEAMSPT 180 SSLTSLREDA DSGDHGQSSI SGTDYIPLFD GDKFRGNDTT YIDGLKGHGI ASWDNVLQCT 240 TELNTDPSLV SFPSILSGSM RNILEQEHNI LDDLLMSKSG PSDEAGSSQS LQSNWQIPFE 300 GNVGHIQALT QSLSSEFGSD YGSGLLRNEA HNGSSEVFSV LSHFHGQPKQ QLMQQNYPEQ 360 HFEGQPQHAL TSDSANRVPG EETINYPLTV RRTLLDRDES LKKVDSFSRW ITKELGEVDD 420 LNMQSSPGIS WSTDECGHVL DDTSLSPSIS QDQLFSINDF SPKWAYAESE IEVLIIGSFL 480 MSQPEVRTSN WSCMFGEVEV HAEVLSNGIL CCQAPHHKVG RVPFYVTCSN RLACSEVREF 540 DYREGFSRKV GIEDFFNSST DMLLHLRLEE LLSLMPVHPP NQTFEGDVEK RNLIFNLISL 600 REEEEFSTKE ELTVEMDISQ QKEHLFCRQV KEKLYSWLLH KVTEGGKGPN VLDNDGQGAL 660 HLAAVLGYDW AITPILTAGV NINFRDVNGW TALHWAASCG RERTVAFLVS MGADSGALTD 720 PSPAFPLGRT AADLASSNGH KGISGFLAES SLTSHLESLT MDDQLKGGRQ EISGMKAVQT 780 VSERTPTPVL YNDMPDALCL KDSLTAVRNA TQAADRIHQV FRMQSFQRKQ LTQTEDGEFD 840 LLDQQALSLL ASKASKSGLG DGLANAAAVQ IQKKFRGWKK RKEFLIIRQR IVKIQAHVRG 900 HQVRKQYKTV IWSVGILEKV ILRWRRKGSG LRGFRPDAVN KVPCLQQNDS QKEDEYDYLK 960 EGRKQSEEKI QKALSRVKSM VQYPEARAQY RRLLNVVDDF RQKKASNMDL IHSEETVDGM 1020 EDLIDIDMLL DDDNFIPIAF D |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, old leaves, petals, sepals, top of carpels, stigmas, stamen filaments, anthers and siliques, but not in pollen. {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:14581622}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
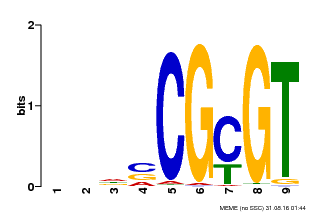
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lj6g3v2218220.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003593198.2 | 0.0 | calmodulin-binding transcription activator 2 isoform X2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | G7INW0 | 0.0 | G7INW0_MEDTR; Calmodulin-binding transcription activator 1 | ||||
| STRING | AES63449 | 0.0 | (Medicago truncatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7406 | 26 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



