 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lj6g3v0528550.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Loteae; Lotus
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1106aa MW: 125104 Da PI: 5.7493 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 173.1 | 3.8e-54 | 21 | 136 | 3 | 118 |
CG-1 3 kekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqr 96
+ ++rwl++ ei++iL n++++++++e++ p+sgsl+L++rk++ryfrkDG++w+kkkdgktvrE+he+LK g+v+vl+cyYah+e +++fqr
Lj6g3v0528550.1 21 EAQHRWLRPAEICEILSNYRNFRIASEPAYMPPSGSLFLFDRKVMRYFRKDGHNWRKKKDGKTVREAHERLKAGSVDVLHCYYAHGEGHENFQR 114
669******************************************************************************************* PP
CG-1 97 rcywlLeeelekivlvhylevk 118
r+yw+Leeel++iv+vhy++vk
Lj6g3v0528550.1 115 RTYWMLEEELSHIVFVHYRDVK 136
*******************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 76.695 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 9.4E-74 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 5.5E-48 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF01833 | 1.1E-6 | 521 | 597 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 3.11E-17 | 521 | 606 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:2.60.40.10 | 1.2E-5 | 521 | 595 | IPR013783 | Immunoglobulin-like fold |
| PROSITE profile | PS50297 | 17.024 | 706 | 817 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 9.64E-17 | 712 | 817 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 3.8E-17 | 714 | 821 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 9.79E-13 | 717 | 815 | No hit | No description |
| Pfam | PF12796 | 3.6E-6 | 728 | 818 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 8.656 | 756 | 788 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.038 | 756 | 785 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 1400 | 795 | 824 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 7.43E-7 | 925 | 980 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.26 | 930 | 952 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.133 | 931 | 960 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0078 | 933 | 951 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.069 | 953 | 975 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.041 | 954 | 978 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 7.1E-4 | 956 | 974 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1106 aa Download sequence Send to blast |
MAETTRYVPP NQLDIEQIIL EAQHRWLRPA EICEILSNYR NFRIASEPAY MPPSGSLFLF 60 DRKVMRYFRK DGHNWRKKKD GKTVREAHER LKAGSVDVLH CYYAHGEGHE NFQRRTYWML 120 EEELSHIVFV HYRDVKGTKA NYRWAKENEE SLPSQQTYKI MPNTETEVKT SISSTLHPRG 180 YQVPSQTVDT SMNSAQTSEY EEAESAFNNH ANSEFYSFLE LQHPVVENSK AQIADSYCPL 240 PLKDDRAKFP VIPGVNYTSL GQANKIKDIH NVGLTYEPSK SLGFSSWEDI LGNNGGSHHV 300 PFQPSFSETQ SNDKGINRDP SQAYEIMGQH FTISIDEQHG NGSPITPEGS WQASEFNSLS 360 MSNSPIDGLY SGSTCEVIYS NCAQEVYEVD FQKSMEQCLL HPYKQNKFLM QNDPQENSLK 420 EKDSLNSDIE ANRSLDGIED TSLSLKTNLL DGTQEGLKKL DSFNQWMSKE LGDVEESNKQ 480 STSDAYWDAV ESENGVDSTT IPSLDTYVLD PSVSHDQLFS IIDYSPSWTF EGSETMVLIS 540 GQFLRSQHEA EQCKWSCMFG EKEVPAEIIE NGVLCCHTPP HKAGRVPFYV TCSNRLACSE 600 VREFDFRVNY SPEINTAGEN RGSIVDIYNI RFGELLSLEH ALPSSSDSQS ISEKSELRNE 660 ISSLLRREDD DWDKLLKLTL EIDFSPEILQ EQLLQNLLKD KLLAWLLHKI TDDGKGPNVL 720 DGGGQGVLHF AAALGYDWAL EPTVVAGVNV NFRDVHGWTA LHWAAFCGRE RTVASLISLG 780 AAPVALTDPC LAYPSGRTPA DLASANGHKG IAGFLAQSFL HAQLSSLDLN RNMGETSAAK 840 VFHRIQEQNT AQVNHYEGLS YELSLKDSLA AVCNATQAAA RIHQVFRVQS FQRKQLKEHG 900 EDRFGISDEQ ALSLVTKNVK PHKSGQRNEP AHVAATRIQN KFRGWKGRKE FLLIRQRIVK 960 IQAHVRGHQV RKNHGKIIWS VGILEKVVLR WRRKGCGLRG FKSEGVSEGN MVQGESSTED 1020 EYDLLKEGRK QTEQRLQIAL ARVKSMVQYP EARDQYHRLL NVVTEIQENQ VKHERTSNNS 1080 EETREFDNLI DLEALLDEDI FMPTAT |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Lja.20415 | 0.0 | root | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
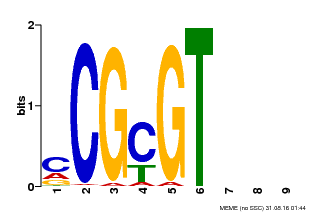
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lj6g3v0528550.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003524262.1 | 0.0 | calmodulin-binding transcription activator 1-like isoform X2 | ||||
| TrEMBL | I1K4J1 | 0.0 | I1K4J1_SOYBN; Uncharacterized protein | ||||
| STRING | GLYMA05G31190.2 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8284 | 27 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



