 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lj2g3v1989250.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Loteae; Lotus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 304aa MW: 33856.9 Da PI: 7.1293 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.4 | 1.2e-18 | 140 | 194 | 2 | 56 |
T--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 2 rkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
rk+ +++k+q +Lee F+++++++ +++ LAk+lgL rqV vWFqNrRa+ k
Lj2g3v1989250.1 140 RKKLRLSKDQAIVLEESFKEHNTLNPKQKMALAKQLGLRARQVEVWFQNRRARTK 194
788899***********************************************98 PP
| |||||||
| 2 | HD-ZIP_I/II | 125.8 | 1.9e-40 | 140 | 229 | 1 | 91 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreel 91
+kk+rlsk+q+ +LEesF+e+++L+p++K++la++Lgl++rqv+vWFqnrRARtk+kq+E+d+e+Lkr++++l+een+rL+kev+eLr +l
Lj2g3v1989250.1 140 RKKLRLSKDQAIVLEESFKEHNTLNPKQKMALAKQLGLRARQVEVWFQNRRARTKLKQTEVDCEFLKRCCENLTEENRRLQKEVQELR-AL 229
69*************************************************************************************9.55 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF04618 | 3.7E-27 | 4 | 111 | IPR006712 | HD-ZIP protein, N-terminal |
| Gene3D | G3DSA:1.10.10.60 | 2.6E-18 | 125 | 194 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 3.51E-18 | 131 | 197 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.297 | 136 | 196 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 6.2E-15 | 138 | 200 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 8.83E-15 | 140 | 197 | No hit | No description |
| Pfam | PF00046 | 4.7E-16 | 140 | 194 | IPR001356 | Homeobox domain |
| PRINTS | PR00031 | 1.9E-5 | 167 | 176 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 171 | 194 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.9E-5 | 176 | 192 | IPR000047 | Helix-turn-helix motif |
| CDD | cd14686 | 0.0067 | 189 | 228 | No hit | No description |
| Pfam | PF02183 | 2.3E-11 | 196 | 230 | IPR003106 | Leucine zipper, homeobox-associated |
| SMART | SM00340 | 4.2E-27 | 196 | 239 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009641 | Biological Process | shade avoidance | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 304 aa Download sequence Send to blast |
MTLGKEDDLG LSLSLSLSFP QKNPPNPNPL HLNLMSSSSA YNPLKPTWTQ PFTSSDRDSE 60 TCRVGEGGFS LGGIGIDVNR LPSGVDCEEE AGVSSPNSTV SSVSGKRSER EGNGEERDVE 120 ERDCSRGISD EDDDGGETSR KKLRLSKDQA IVLEESFKEH NTLNPKQKMA LAKQLGLRAR 180 QVEVWFQNRR ARTKLKQTEV DCEFLKRCCE NLTEENRRLQ KEVQELRALK LSPQFYMQMT 240 PPTTLTMCPS CERVAVPPSA VDPSTRHHHP MSAPNNHARP FSVGAWASTA PISHRPFDVL 300 RPRS |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 138 | 144 | SRKKLRL |
| 2 | 188 | 196 | RRARTKLKQ |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Lja.9605 | 0.0 | cell culture| pod| root | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that plays a role in auxin-mediated morphogenesis. Negatively regulates lateral root elongation. {ECO:0000269|PubMed:12492842}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00550 | DAP | Transfer from AT5G47370 | Download |
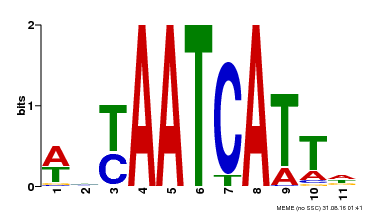
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lj2g3v1989250.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By indole-3-acetic acid (IAA). {ECO:0000269|PubMed:12492842}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003611347.1 | 1e-159 | homeobox-leucine zipper protein HAT4 | ||||
| Swissprot | P46601 | 1e-108 | HAT2_ARATH; Homeobox-leucine zipper protein HAT2 | ||||
| TrEMBL | B7FJ62 | 1e-157 | B7FJ62_MEDTR; Homeobox associated leucine zipper protein | ||||
| STRING | AES94305 | 1e-158 | (Medicago truncatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1239 | 34 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16780.1 | 4e-95 | homeobox protein 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



