|
Plant Transcription
Factor Database
|
Transcription Factor Information
|
Basic
Information? help
Back to Top |
| TF ID |
Lj2g3v1728900.1 |
| Organism |
|
| Taxonomic ID |
|
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Loteae; Lotus
|
| Family |
SRS |
| Protein Properties |
Length: 321aa MW: 33991.2 Da PI: 7.0123 |
| Description |
SRS family protein |
| Gene Model |
| Gene Model ID |
Type |
Source |
Coding Sequence |
| Lj2g3v1728900.1 | genome | Kazusa | View CDS |
|
| Signature Domain? help Back to Top |
 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | DUF702 | 204.9 | 2.2e-63 | 109 | 255 | 3 | 153 |
DUF702 3 sgtasCqdCGnqakkdCaheRCRtCCksrgfdCathvkstWvpaakrrerqqqlaaasskaaasaaeaaskrkrelkskkqsalsstkls.sae 95
g+asCqdCGnqakkdC h RCRtCCksrgfdC+th+kstWvpa+krrerqqq++++ + + ++++ ++ + + ++s+l++t l+ + +
Lj2g3v1728900.1 109 GGGASCQDCGNQAKKDCPHVRCRTCCKSRGFDCQTHLKSTWVPASKRRERQQQQQQQLVPTRQ-QRDHGHNVH----NPTSSSLACTLLPpNPS 197
5789**********************************************9988876544433.222222222....33566777777761445 PP
DUF702 96 skkeletsslPeevsseavfrcvrvssvddgeeelaYqtavsigGhvfkGiLydqGle 153
++ le ++P+ vs++a f+cvrvssvd+++ee+aY tav+igGhvf+GiLyd+G++
Lj2g3v1728900.1 198 HSGGLEEVNFPAVVSTPAEFKCVRVSSVDEADEEYAYSTAVNIGGHVFRGILYDHGPD 255
556788899***********************************************98 PP
|
| Sequence ? help Back to Top |
| Protein Sequence Length: 321 aa
Download sequence Send
to blast |
MAGLFSLGGG GGRGNDAEDQ QHHHQQNEIP PPPDSSLFWY SNKNHDDVVS SSYRGFEIWN 60
QQQQHLVPAQ PQPRAFFQQA DLYTSAAALG VGPSEASRSG KSGGSGGGGG GASCQDCGNQ 120
AKKDCPHVRC RTCCKSRGFD CQTHLKSTWV PASKRRERQQ QQQQQLVPTR QQRDHGHNVH 180
NPTSSSLACT LLPPNPSHSG GLEEVNFPAV VSTPAEFKCV RVSSVDEADE EYAYSTAVNI 240
GGHVFRGILY DHGPDGNSYT AAGAGSSSST GGGVVQPLNL ISAAGNTTAV GAIVDPSSLY 300
PAPLNAFMPA SGTQFFPHPR P
|
| Expression --
Description ? help
Back to Top |
| Source |
Description |
| Uniprot | DEVELOPMENTAL STAGE: Expressed in the apical parts of the developing gynoecium. Detected throughout the youngest flower primordium. Later relocalizes towards the regions of the presumptive sepal anlagen and remains in sepal primordia until just after their emergence. Also observed on the abaxial side of the young floral meristem. Present in the newly arisen gynoecial primordium. Restricted to the apical parts of the carpels as the open-ended gynoecial cylinder elongates vertically. In the apical regions of the gynoecium, confined to a zone in the interphase between the style and the stigma and fades out later. Within the gynoecium, accumulates in ovule primordia, and, as the ovules developed, restricted to the epidermis of the developing funiculi, to the outer, but not the inner, integuments and to the tip of the nucellus. Also present in the cell layer of the septum that faces the ovary. In the embryo, detected in the cotyledon primordia during late globular to mid heart stage. In addition, transiently expressed in petal and stamen primordia and in the tapetum of the anthers. {ECO:0000269|PubMed:12361963}. |
| Uniprot | DEVELOPMENTAL STAGE: In germinating seeds, first detected at about 48 h after imbibition. In young seedlings grown in continuous light, present in the root/shoot transition zone, in the apex, and in the apical region of the cotyledons. Later observed in lateral root primordia and in emerging lateral roots, particularly in the root tips, as well as in the shoot apex and in the new leaves, especially in the apical and lateral hydathodes. In adult plants, accumulates in lateral root tips, lateral root primordia, and axillary shoot primordia. In flowers, expressed in the style and stigmatic surface of the pistil and in the receptacle (base) of the flower throughout the development of the pistil until anthesis and later in the silique. Fades out after fertilization. {ECO:0000269|PubMed:11706176, ECO:0000269|PubMed:16740146}. |
| Uniprot | TISSUE SPECIFICITY: Expressed in flowers, seeds and seedlings. {ECO:0000269|PubMed:12361963}. |
| Uniprot | TISSUE SPECIFICITY: Expressed in young organs such as shoot apices and root tips. Present in roots, stems, flowers, leaves, hydathodes and siliques. {ECO:0000269|PubMed:10368174, ECO:0000269|PubMed:11706176}. |
| Functional Description ? help
Back to Top |
| Source |
Description |
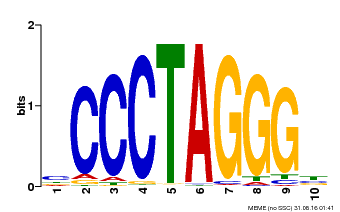
| UniProt | Transcription activator that binds DNA on 5'-ACTCTAC-3' and promotes auxin homeostasis-regulating gene expression (e.g. YUC genes), as well as genes affecting stamen development, cell expansion and timing of flowering. Synergistically with other SHI-related proteins, regulates gynoecium, stamen and leaf development in a dose-dependent manner, controlling apical-basal patterning. Promotes style and stigma formation, and influences vascular development during gynoecium development. May also have a role in the formation and/or maintenance of the shoot apical meristem (SAM). {ECO:0000269|PubMed:12361963, ECO:0000269|PubMed:16740145, ECO:0000269|PubMed:16740146, ECO:0000269|PubMed:18811619, ECO:0000269|PubMed:20154152, ECO:0000269|PubMed:22318676}. |
| UniProt | Transcription activator that binds DNA on 5'-ACTCTAC-3' and promotes auxin homeostasis-regulating gene expression (e.g. YUC genes), as well as genes affecting stamen development, cell expansion and timing of flowering. Synergistically with other SHI-related proteins, regulates gynoecium, stamen and leaf development in a dose-dependent manner, controlling apical-basal patterning. Promotes style and stigma formation, and influences vascular development during gynoecium development. May also have a role in the formation and/or maintenance of the shoot apical meristem (SAM). Suppressor of the gibberellin (GA) signal transduction. {ECO:0000269|PubMed:10368174, ECO:0000269|PubMed:11706176, ECO:0000269|PubMed:16740146}. |
| Regulation -- Description ? help
Back to Top |
| Source |
Description |
| UniProt | INDUCTION: Regulated by ESR1 and ESR2. {ECO:0000269|PubMed:21976484}. |
| Annotation --
Nucleotide ? help
Back to Top |
| Source |
Hit ID |
E-value |
Description |
| GenBank | AP006643 | 0.0 | AP006643.1 Lotus japonicus genomic DNA, chromosome 2, clone: LjT34A19, TM0249, complete sequence. |
| GenBank | AP006712 | 0.0 | AP006712.1 Lotus japonicus genomic DNA, chromosome 2, clone: LjT17E09, TM0627, complete sequence. |
| Best hit in Arabidopsis thaliana ? help
Back to Top |
| Hit ID |
E-value |
Description |
| AT3G51060.1 | 4e-63 | Lateral root primordium (LRP) protein-related |
| Publications
? help Back to Top |
- Islam MA, et al.
Overexpression of the AtSHI gene in poinsettia, Euphorbia pulcherrima, results in compact plants.
PLoS ONE, 2013. 8(1): p. e53377
[PMID:23308204] - Pfannebecker KC,Lange M,Rupp O,Becker A
An Evolutionary Framework for Carpel Developmental Control Genes.
Mol. Biol. Evol., 2017. 34(2): p. 330-348
[PMID:28049761] - Estornell LH,Landberg K,Cierlik I,Sundberg E
SHI/STY Genes Affect Pre- and Post-meiotic Anther Processes in Auxin Sensing Domains in Arabidopsis.
Front Plant Sci, 2018. 9: p. 150
[PMID:29491878]
|