 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lj1g3v2611340.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Loteae; Lotus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 810aa MW: 88258 Da PI: 6.216 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 65.6 | 6.6e-21 | 117 | 172 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l L++rqVk+WFqNrR+++k
Lj1g3v2611340.1 117 KKRYHRHTPQQIQELESLFKECPHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 172
688999***********************************************999 PP
| |||||||
| 2 | START | 206.9 | 7.7e-65 | 325 | 548 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE..EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEE CS
START 1 elaeeaaqelvkkalaeepgWvkss..esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetle 83
ela++a++elvk+a+ +ep+Wv++ e++n de+ +++++ + + +ea+r++g+v+ ++ lve+l+d++ +W+e+++ + +t+e
Lj1g3v2611340.1 325 ELALAAMEELVKMAQTGEPLWVLEGgrEILNHDEYNRTVTPCIGlrpngFVSEASRETGMVIINSLALVETLMDSN-RWSEMFPcvvaRTSTTE 417
5899********************999*************9999********************************.***************** PP
EECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--..-TTSEE-EESSEEEEEEEECTCEEEEEEEE- CS
START 84 vissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe.sssvvRaellpSgiliepksnghskvtwveh 169
vis+g galqlm aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+d ++ ++ ++ + +++lpSg+++++++ng+skvtwveh
Lj1g3v2611340.1 418 VISNGingtrnGALQLMHAELQVLSPLVPvREVNFLRFCKQHAEGVWAVVDVSIDTIRETSSgTPTFLNCRRLPSGCVVQDMPNGYSKVTWVEH 511
************************************************************99******************************** PP
EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 170 vdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
++++++++h+l+r+l++sg+ +ga++wva lqrqce+
Lj1g3v2611340.1 512 AEYDESQVHQLYRPLLSSGIGFGAQRWVAILQRQCEC 548
***********************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.03E-20 | 100 | 174 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.9E-21 | 104 | 174 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.443 | 114 | 174 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.2E-17 | 115 | 178 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.82E-18 | 116 | 174 | No hit | No description |
| Pfam | PF00046 | 1.9E-18 | 117 | 172 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 149 | 172 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 44.647 | 316 | 551 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.06E-32 | 318 | 548 | No hit | No description |
| CDD | cd08875 | 6.33E-121 | 320 | 547 | No hit | No description |
| SMART | SM00234 | 4.8E-46 | 325 | 548 | IPR002913 | START domain |
| Pfam | PF01852 | 2.8E-56 | 325 | 548 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.88E-22 | 576 | 803 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 810 aa Download sequence Send to blast |
MSFGGVFENK QGGEGAKIVA DIAYNQDKMP SGAISQPRLA TPTLAKSMLN SSSLSLALQT 60 NIDGQGNGHG NGLIHENYEG HGLRRSREEE HESRSGSDNM DAASGDDFDA ADNRPRKKRY 120 HRHTPQQIQE LESLFKECPH PDEKQRLELS KRLCLETRQV KFWFQNRRTQ MKTQLERHEN 180 SLLRQENDKL RAENMSIREA MRNPMCSNCG GPAVMGEISL EEQHLRIENA RLKDELDRVC 240 ALAGKFLGRP IGPPLLNSSL ELGVGSNGFG GLSNMPSTLG PDFVGISSPL GMVTPPPPPA 300 QSTRTTTAMM SSGFDRSVER SMLLELALAA MEELVKMAQT GEPLWVLEGG REILNHDEYN 360 RTVTPCIGLR PNGFVSEASR ETGMVIINSL ALVETLMDSN RWSEMFPCVV ARTSTTEVIS 420 NGINGTRNGA LQLMHAELQV LSPLVPVREV NFLRFCKQHA EGVWAVVDVS IDTIRETSSG 480 TPTFLNCRRL PSGCVVQDMP NGYSKVTWVE HAEYDESQVH QLYRPLLSSG IGFGAQRWVA 540 ILQRQCECLA ILMSSALPSR EHAAISAGGR RSMLKLAQRM TNNFCAGVCA STVHQWNKLN 600 NTSNNNAGAE EVRVMTRKSV DDPGEPPGIV LSAATSVWLP VAPQRVFDFL RDERLRSEWD 660 ILSNGGPMQE MAHIAKGQDH ANCVSLLRAS AMNANQSSML ILQETCTDAS GSLVVYAPVD 720 IPAMHVVMNG GDSAYVALLP SGFSVVPDGN GQGGGDDRGS ASGGSLLTVA FQILVNSLPT 780 AKLTVESVET VNNLILCTVQ KIKAALNGES |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Lja.10106 | 2e-37 | pod | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves and floral buds. {ECO:0000269|PubMed:10402424}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
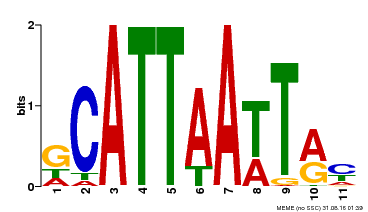
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lj1g3v2611340.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027349905.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A0S3RT53 | 0.0 | A0A0S3RT53_PHAAN; Uncharacterized protein | ||||
| TrEMBL | A0A4D6NHU6 | 0.0 | A0A4D6NHU6_VIGUN; Homeobox-leucine zipper protein | ||||
| STRING | XP_007139955.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1192 | 34 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



