 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lj0g3v0149769.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Loteae; Lotus
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 668aa MW: 73295.1 Da PI: 5.9935 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 86.2 | 3.2e-27 | 214 | 267 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
+pr++W+ eLH++F+ av++L G++kA+Pk+ile+m+v+gLt+e+v+SHLQkYRl
Lj0g3v0149769.1 214 RPRVVWSVELHQQFMAAVNNL-GIDKAVPKKILEIMNVPGLTRENVASHLQKYRL 267
59*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 71.9 | 2.7e-24 | 37 | 145 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHT CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealka 92
vl+vdD+p+ + +l+++l+ y ev+ ++ +e al ll+ ++ +D++l D+ mp+mdG++ll++i e +lp+i+++a + ++ + + +
Lj0g3v0149769.1 37 VLVVDDDPTCLVILERMLRACLY-EVTKCKRAEVALSLLRGNKngFDIVLSDVHMPDMDGFKLLEHIGLEM-DLPVIMMSADDGKDVVMKGVTH 128
89*****************9999.***************777778**********************6654.8********************* PP
TESEEEESS--HHHHHH CS
Response_reg 93 GakdflsKpfdpeelvk 109
Ga d+l Kp+ +e+l +
Lj0g3v0149769.1 129 GACDYLIKPVRIEALKN 145
*************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 3.2E-165 | 12 | 663 | IPR017053 | Response regulator B-type, plant |
| Gene3D | G3DSA:3.40.50.2300 | 2.0E-40 | 34 | 158 | No hit | No description |
| SuperFamily | SSF52172 | 3.98E-34 | 34 | 154 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 1.3E-28 | 35 | 147 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 41.193 | 36 | 151 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 1.4E-21 | 37 | 145 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 1.95E-26 | 38 | 150 | No hit | No description |
| PROSITE profile | PS51294 | 9.819 | 211 | 270 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 6.45E-19 | 212 | 271 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-28 | 213 | 272 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.1E-23 | 215 | 267 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.0E-6 | 216 | 266 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 668 aa Download sequence Send to blast |
MNAKGPIMST PTSSSASKSS DTAAVTAADQ FPAGLRVLVV DDDPTCLVIL ERMLRACLYE 60 VTKCKRAEVA LSLLRGNKNG FDIVLSDVHM PDMDGFKLLE HIGLEMDLPV IMMSADDGKD 120 VVMKGVTHGA CDYLIKPVRI EALKNIWQHV VRKRKNDGKD PDQSGGGDEG DQQLKGSDDA 180 DYSSSANECK STKKRRDQED EDAEDRDDSS SLKRPRVVWS VELHQQFMAA VNNLGIDKAV 240 PKKILEIMNV PGLTRENVAS HLQKYRLYLR RVSGVSQPQN NLNSSFISPY ESAYNASSIN 300 GIDLQTISAH SLAKLQAGLG RSTAKVGVPM PLVEQKNLFS FENSKLRFGE GQLQHMSSNK 360 PMNLLHGIPT NMEPKQFAHW HQSTQSLGNL NMRFNASAAQ RNPFLMQMNH SQPRGQMLGE 420 NTGSHITRFP SSLVQPSGPN GISNGAIENG ITDTNNITPQ RSSLLSFPMN QTTDISASSF 480 PLRSTPGISN ITTKGMFHEE VTLGTKGSGG FFPSYDIFNE LHQKSHDWDP TNSGLTYDAS 540 QHTNPLQGSF DVLPSILGHQ DFSSAQQTAE NRDTTLIGKG MVSMGDSMNL GNLQNVGQHH 600 LNTSLIDNSA RVKAEKFPDP SSQNNLFLEQ YGQDDLLSEL LKQQEGIGVG PIENEFDFDG 660 YSMDNIPV |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 2e-22 | 210 | 273 | 1 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Lja.25468 | 0.0 | pod | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Detected in the whole plant. Predominantly expressed in pollen. {ECO:0000269|PubMed:11370868, ECO:0000269|PubMed:15173562, ECO:0000269|PubMed:9891419}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]GATT-3'. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. Could directly activate some type-A response regulators in response to cytokinins. Involved in the expression of nuclear genes for components of mitochondrial complex I. Promotes cytokinin-mediated leaf longevity. Involved in the ethylene signaling pathway in an ETR1-dependent manner and in the cytokinin signaling pathway. {ECO:0000269|PubMed:11370868, ECO:0000269|PubMed:11574878, ECO:0000269|PubMed:15282545, ECO:0000269|PubMed:16407152}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
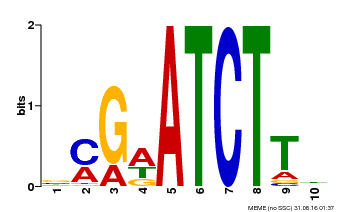
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lj0g3v0149769.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027366964.1 | 0.0 | two-component response regulator ARR2-like | ||||
| Swissprot | Q9ZWJ9 | 0.0 | ARR2_ARATH; Two-component response regulator ARR2 | ||||
| TrEMBL | A0A445GTV5 | 0.0 | A0A445GTV5_GLYSO; Two-component response regulator | ||||
| TrEMBL | I1MGI9 | 0.0 | I1MGI9_SOYBN; Two-component response regulator | ||||
| STRING | GLYMA15G15520.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3958 | 34 | 58 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 1e-170 | response regulator 2 | ||||



