 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lj0g3v0050029.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Loteae; Lotus
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 583aa MW: 65393.1 Da PI: 5.063 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 76.7 | 3e-24 | 200 | 253 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
k+r++W+ +LH++Fv+av+q+ G +k Pk+il+lm+v+ Lt+e+v+SHLQkYRl
Lj0g3v0050029.1 200 KARVVWSVDLHQKFVRAVNQI-GFDKVGPKKILDLMNVPWLTRENVASHLQKYRL 253
68*******************.9*******************************8 PP
| |||||||
| 2 | Response_reg | 80.8 | 4.5e-27 | 20 | 128 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHT CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealka 92
vl+vdD+p+ +++l+++l+k +y +v+++ ++ al ll+e++ +D+++ D++mp+mdG++ll++ e +lp+i+++ ge + + + ++
Lj0g3v0050029.1 20 VLVVDDDPTWLKILEKMLKKCNY-DVTTCCLARHALSLLRERKdgYDIVISDVNMPDMDGFKLLEHVGLEM-DLPVIMMSVDGETSRVMKGVQH 111
89*********************.***************999999**********************6644.8********************* PP
TESEEEESS--HHHHHH CS
Response_reg 93 GakdflsKpfdpeelvk 109
Ga d+l Kp+ ++el++
Lj0g3v0050029.1 112 GACDYLLKPIRMKELRN 128
**************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 2.9E-146 | 2 | 556 | IPR017053 | Response regulator B-type, plant |
| Gene3D | G3DSA:3.40.50.2300 | 4.8E-44 | 17 | 160 | No hit | No description |
| SuperFamily | SSF52172 | 1.27E-35 | 17 | 145 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 3.3E-31 | 18 | 130 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 42.629 | 19 | 134 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 3.6E-24 | 20 | 129 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 4.54E-29 | 21 | 134 | No hit | No description |
| SuperFamily | SSF46689 | 3.23E-17 | 198 | 257 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 9.6E-28 | 198 | 258 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.6E-21 | 200 | 253 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.4E-7 | 202 | 252 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000160 | Biological Process | phosphorelay signal transduction system | ||||
| GO:0009735 | Biological Process | response to cytokinin | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 583 aa Download sequence Send to blast |
MENGCFPSPP RRDFPAGLRV LVVDDDPTWL KILEKMLKKC NYDVTTCCLA RHALSLLRER 60 KDGYDIVISD VNMPDMDGFK LLEHVGLEMD LPVIMMSVDG ETSRVMKGVQ HGACDYLLKP 120 IRMKELRNIW QHVLRKRIHE AREFESFESF EGNYLMRNGS ELSDDGNLFA LEDMTSSKKR 180 KDADNKHDDK EFVDPSSTKK ARVVWSVDLH QKFVRAVNQI GFDKVGPKKI LDLMNVPWLT 240 RENVASHLQK YRLYLSRLQK DDEQKSSSSG MKISDLPTKE VGSFGLHNSV TKQQNDVSID 300 SCNYPDGALQ IQKVETKSQE GNTKGIVSQS TMEEKGRALT GNNNITDVMR EGLQVGLNQP 360 FALPESKGNC TSFDCAIPTK YSWTEVPGMQ LKEEQKSIVQ LEDSFNQLPP LLGKQHHIQV 420 DQSQSIASIS STPSIMKEDV PACIETKPVF SDYKSSYTSS ISSKVSALDT FPIQPGSLMM 480 SDQPLQPTSA ANLGLKTQGS NLSSISDLEF YQRNLLLGGD APSASLDEDL YSYWIQGECY 540 NMNFGLQSVA MPEFYDPGLI AEVPPHLYDS TDYTVIDQGL FIA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 1e-17 | 196 | 258 | 1 | 63 | ARR10-B |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00010 | PBM | Transfer from AT1G67710 | Download |
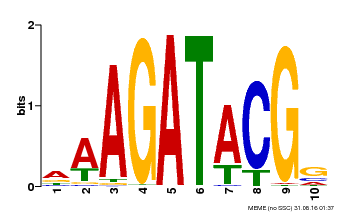
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lj0g3v0050029.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028213070.1 | 0.0 | two-component response regulator ARR11-like isoform X1 | ||||
| TrEMBL | A0A0B2QVM4 | 0.0 | A0A0B2QVM4_GLYSO; Two-component response regulator ARR11 | ||||
| STRING | AES72100 | 0.0 | (Medicago truncatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3999 | 33 | 60 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G67710.1 | 1e-119 | response regulator 11 | ||||



