 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kalax.1988s0002.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 314aa MW: 34198.6 Da PI: 5.5468 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 63.2 | 5.7e-20 | 157 | 207 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+y+GVr+++ +g+++AeIrdp+++g +r++lg+f+ta +Aaka+++a+ kl+g
Kalax.1988s0002.1.p 157 HYRGVRRRP-WGKFAAEIRDPNKRG--ARVWLGTFDTAIDAAKAYDRAAFKLRG 207
7********.**********96655..*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.730.10 | 4.9E-32 | 156 | 216 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.35E-29 | 157 | 215 | No hit | No description |
| PROSITE profile | PS51032 | 24.474 | 157 | 215 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.31E-22 | 157 | 217 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 6.0E-14 | 157 | 207 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 8.0E-37 | 157 | 221 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.8E-13 | 158 | 169 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.8E-13 | 197 | 217 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 314 aa Download sequence Send to blast |
MATPSEVSTL EQIKQHLFGG LLSPLNLKSF GSGSDSSESV VTTSTDSPVS FSNYGAFSKL 60 ESCNSFEFDQ LPNKFFSFET KFDATGFDFA PEAEVIDLVE TSMKKPQIRT SKLSRKPSLN 120 ISLPDKTEWL DLTQSRAAPS AAVASKPEKA VAGDERHYRG VRRRPWGKFA AEIRDPNKRG 180 ARVWLGTFDT AIDAAKAYDR AAFKLRGSKA ILNFPLEIAA SCSAGDAAGV VDCGRKRRIR 240 DEEVEAEVKK VKAEEGVAEE GVAEETVAKP LTPTSWTAFW DFDETEMRGI FNVPPLSPLS 300 PHPALGLAQL MVL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 2e-30 | 156 | 218 | 4 | 66 | ATERF1 |
| 3gcc_A | 2e-30 | 156 | 218 | 4 | 66 | ATERF1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00549 | DAP | Transfer from AT5G47230 | Download |
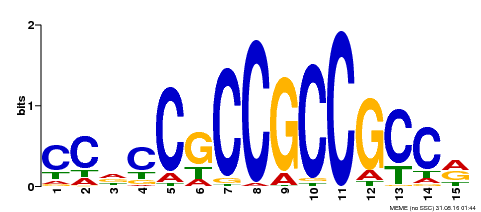
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012070058.1 | 2e-70 | ethylene-responsive transcription factor 5 | ||||
| TrEMBL | A0A067L5Z5 | 4e-69 | A0A067L5Z5_JATCU; Uncharacterized protein | ||||
| STRING | XP_010275423.1 | 1e-68 | (Nelumbo nucifera) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G47230.1 | 5e-41 | ethylene responsive element binding factor 5 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kalax.1988s0002.1.p |



