 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kalax.1507s0001.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 420aa MW: 45475.6 Da PI: 8.0009 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 103.8 | 1e-32 | 229 | 284 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rF++a++qLGG+++AtPk+i+elmkv+gLt+++vkSHLQkYRl+
Kalax.1507s0001.1.p 229 KQRRCWSPELHRRFIHALQQLGGPHVATPKQIKELMKVDGLTNDEVKSHLQKYRLH 284
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-28 | 226 | 287 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 2.15E-18 | 226 | 287 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 15.314 | 226 | 286 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 2.4E-27 | 229 | 284 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 7.0E-8 | 231 | 282 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 420 aa Download sequence Send to blast |
MRRTHLAYNQ STAPALDPAP ARKDRVLWSF YDLGVMDYAD KMQKCSEYVE ALEVERRKIQ 60 VFQRELPLCL ELVSQAIDAC RQQLSSASPP HAAAVGASHE CFHVRQQSEC SEQTSSHGPV 120 LEEFIPIKRA ASCSSEDVGD EEEDESGEKS GDKSGKKSDW LRSVQLWNQS PDPAPVCEVK 180 KNGGAFRPFS KEKAVVVAEM VDKPASSTTL KSRKEESAKE KESGQTQRKQ RRCWSPELHR 240 RFIHALQQLG GPHVATPKQI KELMKVDGLT NDEVKSHLQK YRLHMRRPSA TMASNANAQG 300 PQFVVVGGIW VPPPEYAAPA MTPAATTSSG DPPEADNGKR EARTPSNGVY APVATVGTPF 360 ARPASRTQNS NQKVQSGSHC EDDNGSHSAH GGARAHSGSP ASSSSTHTTA TATASPASS* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in phosphate signaling in roots. {ECO:0000250|UniProtKB:Q9FX67}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
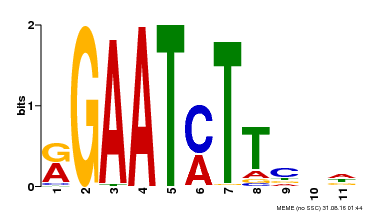
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003631224.1 | 1e-129 | PREDICTED: myb family transcription factor EFM | ||||
| Swissprot | Q9FPE8 | 1e-101 | HHO3_ARATH; Transcription factor HHO3 | ||||
| TrEMBL | A0A438ECN5 | 1e-128 | A0A438ECN5_VITVI; Transcription factor HHO3 | ||||
| STRING | XP_010695844.1 | 1e-125 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G68670.1 | 5e-91 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kalax.1507s0001.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



