 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kalax.0654s0015.3.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 253aa MW: 28572.4 Da PI: 7.5079 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 90.3 | 1.8e-28 | 196 | 247 | 1 | 52 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQk 52
k+r++W+peLH+rFv+a++qLGGse+AtP++i+++m v+gLt+++vkSHLQ
Kalax.0654s0015.3.p 196 KQRRCWSPELHRRFVDALQQLGGSEVATPRQIRDIMRVEGLTNDEVKSHLQV 247
79*************************************************6 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.76E-13 | 194 | 247 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 9.517 | 195 | 252 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.5E-23 | 195 | 248 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.2E-23 | 196 | 249 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.4E-5 | 198 | 248 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 253 aa Download sequence Send to blast |
MDPIPTRDFL KFDDPPRRKL QRLADHVAGL EAERRKIEAF KRELPLCLDL LNDAVERFRE 60 ELRKCGDGGA ADCNCDKVEG PNSENDNDMK SWMSSVKLWN HHANDSPDFN STSSNVKSMI 120 EAEAVAENAG GAKMEIKGNS GDEVSLMKPV HAVARTVELE QLCGESARNL NRGKTLVHEL 180 KMQKKIQQAQ EQVGRKQRRC WSPELHRRFV DALQQLGGSE VATPRQIRDI MRVEGLTNDE 240 VKSHLQVMKT FR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 6e-14 | 196 | 252 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 6e-14 | 196 | 252 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 6e-14 | 196 | 252 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 6e-14 | 196 | 252 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
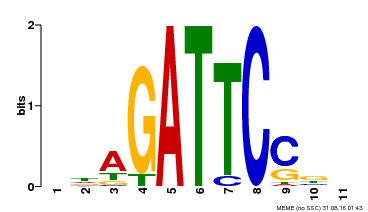
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00629 | PBM | Transfer from PK14402.1 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004288067.1 | 1e-54 | PREDICTED: probable transcription factor KAN4 | ||||
| TrEMBL | A0A2P6RN79 | 9e-49 | A0A2P6RN79_ROSCH; Putative transcription factor MYB-HB-like family | ||||
| STRING | XP_004288067.1 | 4e-54 | (Fragaria vesca) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25550.1 | 2e-43 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kalax.0654s0015.3.p |



