 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kalax.0472s0020.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 977aa MW: 107022 Da PI: 5.814 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 132.3 | 1.6e-41 | 146 | 222 | 1 | 77 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S-- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkq 77
+Cqve+C adls+ak+yhrrhkvCe+hska+ +lv +++qrfCqqCsrfh+l efDe+krsCrrrLa+hn+rrrk++
Kalax.0472s0020.1.p 146 VCQVEDCGADLSKAKDYHRRHKVCEMHSKASRALVGNVMQRFCQQCSRFHALPEFDEGKRSCRRRLAGHNKRRRKTN 222
6**************************************************************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 3.2E-33 | 139 | 208 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.146 | 144 | 221 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 4.97E-38 | 146 | 223 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 6.7E-30 | 147 | 220 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 1.66E-5 | 753 | 860 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.1E-4 | 756 | 859 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 9.64E-5 | 757 | 860 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 977 aa Download sequence Send to blast |
MEANVGDQIR HYYGLGSSSL NPVGKRSLEW DLNDWKWDGH QFVASPLNCK PSGYSSHQLL 60 PETGGSSNNS STCSDDATIG NDTREEELLR SRRAAAGGEV GDAGNLSLML GGNGVPASEM 120 QLTSWEGANG KKSKVVVGGS SSNRGVCQVE DCGADLSKAK DYHRRHKVCE MHSKASRALV 180 GNVMQRFCQQ CSRFHALPEF DEGKRSCRRR LAGHNKRRRK TNPLNVAPGT SLNDDQSNNY 240 LLLTLLKILA DLHSNRSDGA KNQDMLSHLL RSLEENTVAN GGKNITEAVC ALLSNGRQAL 300 ASSGGQHTLA MQEMPQNGSP SADVSPPPST VISIKDSPPG YSVAPSTAAG PVKLNTFDLN 360 DVYVDSADGI EDLEGFRVPA NNGNSSIGCP SWMQQDVHSS SPPQVSRNSD SGSAQSPSSS 420 SGGDQIRTDR IVFKLFGKEP CDFPLALRTQ ILDWLAHTPT DIESYIRPGC IVLTIYLRLA 480 ESVWDELCLD LSSSLSRLID VSEDSFWSTG WVYVRVQDQV AFICNGQVVL DTTSPYISSA 540 SRILSVRPIA IPMSEKASFS IKGCNLFPPN SRLLCAIEGQ YLAQENIPES LEESSSNHND 600 SDKLQVINYS CSIPAITGRG FVEVEEDNGL GRDFFPFIVA ESDLCSEIRM LESTLEPTET 660 DETSENGEPD ARSNAIDFIQ EMGWLLRRSQ LRLRFGYSNK RAEDLFHFDR FKWLMEFSLD 720 RDWCAVVKKL LDILCDGVVG SGEHDSLVAA VSDMGILHRA VRRNSRPLVE LLLRYVPEQT 780 SNNSMMKAGE LLFMPNEMGP AGLTPLHIAA GRDGAEAVLD ALTDDPNKVG IEAWQSARDS 840 TGASPEDYAR LRGHYSYIHL VQRKISKKSF TPGHVVLDIP NTLSDPKANK RETTTQKSSF 900 QISNFPMRTG SGPCKICDQK LIYRTVNTSL AYKPAMLSMV AVAAVCVCVA LLFKSCPEVL 960 YVFHPFRWEN LEYGSS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-30 | 139 | 220 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000269|PubMed:16554053}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
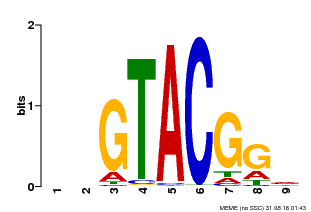
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021664716.1 | 0.0 | squamosa promoter-binding-like protein 1 isoform X2 | ||||
| Swissprot | Q9S7P5 | 0.0 | SPL12_ARATH; Squamosa promoter-binding-like protein 12 | ||||
| TrEMBL | F6HZE3 | 0.0 | F6HZE3_VITVI; Uncharacterized protein | ||||
| STRING | VIT_07s0005g02260.t01 | 0.0 | (Vitis vinifera) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kalax.0472s0020.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



