 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kalax.0301s0019.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 976aa MW: 106837 Da PI: 5.9829 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 131.6 | 2.8e-41 | 145 | 221 | 1 | 77 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S-- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkq 77
+Cqve+C adls+ak+yhrrhkvCe+hska+ +lv ++qrfCqqCsrfh+l efDe+krsCrrrLa+hn+rrrk++
Kalax.0301s0019.1.p 145 VCQVEDCGADLSKAKDYHRRHKVCEMHSKASRALVGTVMQRFCQQCSRFHALPEFDEGKRSCRRRLAGHNKRRRKAN 221
6**************************************************************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 7.3E-33 | 138 | 207 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.026 | 143 | 220 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.83E-37 | 144 | 222 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 1.3E-29 | 146 | 219 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 1.51E-5 | 752 | 859 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.4E-4 | 755 | 858 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 9.63E-5 | 756 | 859 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 976 aa Download sequence Send to blast |
MEANVGDQIR HFYGLGSSSL NPVGKRSLEW DLNDWKWDGH LFVASPLNCK PSGYSSHQLL 60 PETGGSSNNS STCSDDATIG NDTREQELLR SRRAAGGEVG DAGNLSLMLG GNGVPASEMQ 120 LTSWEGANGK KSKVVVGGSS SNRAVCQVED CGADLSKAKD YHRRHKVCEM HSKASRALVG 180 TVMQRFCQQC SRFHALPEFD EGKRSCRRRL AGHNKRRRKA NPLNVAPGTS LNDDQSNNYL 240 LLTLLKILAD LHSNRSDGAK NQDMLSHLLR SLEENTVANG GKNITEAVCA MLSNGRQALA 300 SSGGQHTLAM QEMPQNGSPS ADVSPPPSTV ISIKDSPPGY SVAPSTAAGP VKLNTFDLND 360 VYVDSADGID DLEGFRVPAN NGNSSIGCSS WMQQDVHSSS PPQVSRNSDS GSAQSPSSSS 420 GGNQIRTDRI VFKLFGKEPC DFPLALRTQI LDWLAHTPTD IESYIRPGCI VLSIYLRLAE 480 SVWDELCLDL SSSLSRLIDV SEDSFWSTGW VYVRVQDQVA FICNGQVVLD TTLPYISSAS 540 RILSVRPIAV PMSEKASFSI KGCNIFPPNS RLLCALEGQY LAQENIPESL EESSSNHNDS 600 DKLQVINYSC SIPAIIGRGF VEVEEDNGLG RDFFPFIVAE SDICSEIRML ESTLEPTETD 660 ETSENGEPDA RSNAIDFIQE MGWLLRRSQL RLRFGYSNKR AEDLFHLDRL KWLMEFSLDR 720 DWCAVVKKLL DILCDGVVGS GEHDSLVAAV SDMGILHRAV RRNSRPLVEL LLRYVPEQTS 780 NNSMMKGGEL LFMPNEMGPA GLTPLHIAAG RDGAEAVLDA LTDDPNKVGI EAWQSARDST 840 GASPEDYARL RGHYSYIHLV QRKISKKSFT PGHVVLDIPN TLSDPKANKR ETTTQKSSFQ 900 ILNFPMRTGS GPCKICDQKL IYRTVNTSLA YKPAMLSMVA VAAVCVCVAL LFKSCPEVLY 960 VFHPFRWENL EYGSS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 2e-30 | 138 | 219 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000269|PubMed:16554053}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
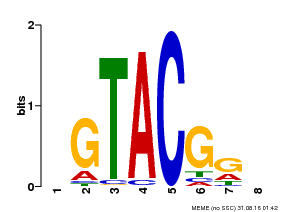
| Motif ID | Method | Source | Motif file |
| MP00060 | PBM | Transfer from AT3G60030 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021664716.1 | 0.0 | squamosa promoter-binding-like protein 1 isoform X2 | ||||
| Swissprot | Q9S7P5 | 0.0 | SPL12_ARATH; Squamosa promoter-binding-like protein 12 | ||||
| TrEMBL | F6HZE3 | 0.0 | F6HZE3_VITVI; Uncharacterized protein | ||||
| STRING | VIT_07s0005g02260.t01 | 0.0 | (Vitis vinifera) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kalax.0301s0019.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



