 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kalax.0168s0032.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 847aa MW: 96886 Da PI: 6.9166 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 82.5 | 7.1e-26 | 93 | 195 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk............tekerrtraetrtgCkaklkvkkekdgkwev 78
+fY+eYA+ GF++ +++s++sk+++e+++++f Cs++g+++e +k+ +++ +r+ +t+Cka+++vk++ dgkw++
Kalax.0168s0032.1.p 93 SFYQEYARANGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYDKSlnkprarqvkqePDNTAGRRSCAKTDCKASMHVKRRPDGKWVI 182
5*******************************************9999999988776544444458999********************* PP
FAR1 79 tkleleHnHelap 91
+++++eHnHel+p
Kalax.0168s0032.1.p 183 HRFVKEHNHELQP 195
***********86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 9.3E-24 | 93 | 195 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 1.0E-26 | 293 | 384 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.087 | 573 | 609 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 2.3E-5 | 581 | 606 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 2.4E-7 | 584 | 611 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 847 aa Download sequence Send to blast |
MDIDLRLPTG KTDREEEEDA NGFESILDGD IKLSNGDGEL GNLGIVGGSD NNIEYEHEGD 60 MNSHEHDMVD YKEDSNLEPI SGMEFESHGE AYSFYQEYAR ANGFNTAIQN SRRSKTSREF 120 IDAKFACSRY GTKREYDKSL NKPRARQVKQ EPDNTAGRRS CAKTDCKASM HVKRRPDGKW 180 VIHRFVKEHN HELQPAQAVS EQTRKMYAAM ARQFAEYKNL SGLKNDSKNP YDKSRGSSIE 240 AGDLRCLLDY CTQMQALNSN FFYSIDLGDD QRLRSLLWVD AKSRHDYTYF NDIVSFDTTY 300 VRSKYRFPLA LFVGVNQHYQ FVLLGCALMF EESPPTFNWI MQAWLRAMGG QAPKVIITDQ 360 DRSLKQVVSD VFPNAAHCFC LWNVLGKVSE NLSHVMKHHE NFMQKFEKCI YRSGTVDEFD 420 RRWGKIVERF ELKEDGFFQS LYEDQKHWVP LYMRDLVLAG MSTPQRSDSV NSSFDKYLHK 480 KTSVQDFIKQ HEAFLQDRYD EEARADSDPW NKLPITKSPS PLEKHMSVAY THAVFKKGQV 540 ELLGAIACHP KPERQEGAEM VFRVQDVEKK QEFAVAWNEL KAEISCVCRL FEYKGFLCRH 600 AMIVLQICGQ SQIPSHYILK RWAKDAKMRP LMPEAPLQAQ SRVHRYDDLC QRAMRLGEEA 660 ALSQESYEIA FRAIDEALSN CAAVNNNSGR SYVEAGMSSA SHGLRVDEEN QSLNKMSKKK 720 NSIKKRKVTP EPDDMTVEAQ NLQQMDKIGV RSVTLDSIYG TQHNVQGMVQ LNLMGPTRDN 780 FYSNPQGLGQ LNSIAPAHDG YYVPPPSMHN LAGQMDFFRT PAGFYGIRDE SSIRATQIHG 840 DHTRHG* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
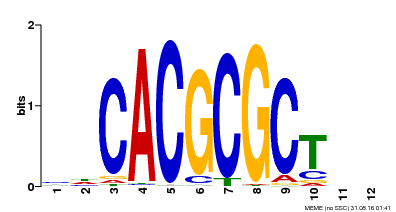
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008229656.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A251Q297 | 0.0 | A0A251Q297_PRUPE; Uncharacterized protein | ||||
| STRING | XP_008229656.1 | 0.0 | (Prunus mume) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kalax.0168s0032.1.p |



