 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kalax.0073s0009.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 494aa MW: 54198.2 Da PI: 6.9453 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.2 | 6.5e-32 | 272 | 327 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+p+LH+rFv+a++ LGGs++AtPk+i+elmkv+gLt+++vkSHLQkYRl+
Kalax.0073s0009.1.p 272 KARRCWSPDLHRRFVNALQMLGGSQVATPKQIRELMKVDGLTNDEVKSHLQKYRLH 327
68****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50873 | 9.444 | 17 | 325 | IPR002016 | Haem peroxidase, plant/fungal/bacterial |
| PROSITE profile | PS51294 | 12.22 | 269 | 329 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.0E-28 | 270 | 330 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 3.91E-17 | 270 | 330 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.4E-26 | 272 | 327 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 8.8E-7 | 274 | 325 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006979 | Biological Process | response to oxidative stress | ||||
| GO:0009266 | Biological Process | response to temperature stimulus | ||||
| GO:0009740 | Biological Process | gibberellic acid mediated signaling pathway | ||||
| GO:0010452 | Biological Process | histone H3-K36 methylation | ||||
| GO:0048579 | Biological Process | negative regulation of long-day photoperiodism, flowering | ||||
| GO:0055114 | Biological Process | oxidation-reduction process | ||||
| GO:1903507 | Biological Process | negative regulation of nucleic acid-templated transcription | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0004601 | Molecular Function | peroxidase activity | ||||
| GO:0020037 | Molecular Function | heme binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 494 aa Download sequence Send to blast |
MTPPSELSLD FKPHSYSMLL KSFASGEHQQ PDLKQLEDFL ACLEDERLKI EAFKRELPLC 60 MQLLSNAVEA SRQKLQAVKS SNQGPRPVLE EFIPLKSPTA DGEEDHAASK NASDKSNWMV 120 SAQLWSQTPS TDNNGLITRQ LESPYQTDMG LSITSSNLTL LDTSHTTKQR NSGAFSPFNK 180 DMTSTGPNLP NPNLALANTN DHHLLEEKKI GSELIPDCPT RETIITPGNF NGAGMMDDEP 240 MKGTNNSADT LTTSNTTHTT NNNSSSNQPH RKARRCWSPD LHRRFVNALQ MLGGSQVATP 300 KQIRELMKVD GLTNDEVKSH LQKYRLHTRR PSPSPQPAAG GPAAQLVVLG GIWVPPEYAA 360 HHSGATAALY GAAPHQAVGP EFYAAAGPHH RQLFHIHQQQ QMQMHQIHLQ QQQQHHQAVQ 420 VYNNKASSRT HSSPESDARG AVGDRSESIE DGKSESGSWK GGESVENGRE RMKVGARRDG 480 GEESNGSEIT LKF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 1e-13 | 272 | 325 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 1e-13 | 272 | 325 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 9e-14 | 272 | 325 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 9e-14 | 272 | 325 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 9e-14 | 272 | 325 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 9e-14 | 272 | 325 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 1e-13 | 272 | 325 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 1e-13 | 272 | 325 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 1e-13 | 272 | 325 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 1e-13 | 272 | 325 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 1e-13 | 272 | 325 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 1e-13 | 272 | 325 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00261 | DAP | Transfer from AT2G03500 | Download |
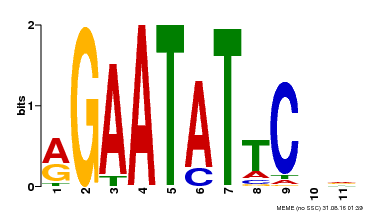
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010098958.1 | 1e-150 | myb family transcription factor EFM | ||||
| TrEMBL | W9R8R3 | 1e-149 | W9R8R3_9ROSA; Two-component response regulator | ||||
| STRING | XP_010098958.1 | 1e-150 | (Morus notabilis) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G03500.1 | 6e-83 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kalax.0073s0009.1.p |



