 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kalax.0039s0143.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 372aa MW: 41053.2 Da PI: 8.5789 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 44.3 | 4.4e-14 | 63 | 111 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s++kGV + +grW A+I++ ++r++lg+f +eeAa+a++ a+++++g
Kalax.0039s0143.1.p 63 SKFKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFNEEEEAARAYDTAAQRFRG 111
79****9888.8*********3.....5*************************98 PP
| |||||||
| 2 | B3 | 97.4 | 8.8e-31 | 190 | 292 | 1 | 94 |
EEEE-..-HHHHTT-EE--HHH.HTT.........---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.........ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvv 81
f+kv tpsdv+k++rlv+pk++ae+h + +++ l +ed g++W+++++y+++s++yvltkGW++Fvk++gL +gD v
Kalax.0039s0143.1.p 190 FEKVVTPSDVGKLNRLVIPKQHAEKHfplntspsaSDAASKGVLLNFEDGGGKVWRFRYSYWNSSQSYVLTKGWSRFVKEKGLLAGDAVS 279
99*************************9999988855556999*********************************************** PP
EEE-SSSEE..EE CS
B3 82 Fkldgrsefelvv 94
F++++ +++l++
Kalax.0039s0143.1.p 280 FERSNGADRQLYI 292
**87666666665 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 5.1E-9 | 63 | 111 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 8.75E-26 | 63 | 118 | No hit | No description |
| SuperFamily | SSF54171 | 1.77E-17 | 63 | 119 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 2.5E-29 | 64 | 125 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 20.152 | 64 | 119 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 7.1E-21 | 64 | 119 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 1.4E-38 | 185 | 297 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 3.14E-30 | 185 | 292 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 5.18E-28 | 189 | 285 | No hit | No description |
| Pfam | PF02362 | 1.6E-28 | 190 | 293 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 3.1E-24 | 190 | 298 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 14.325 | 190 | 298 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 372 aa Download sequence Send to blast |
MDGSCIDESS ATSDSTSAAV DMVVPRKSAT ETTEKLTRMG SGASVITDSA IGVEAESRKL 60 PSSKFKGVVP QPNGRWGAQI YEKHQRVWLG TFNEEEEAAR AYDTAAQRFR GRDAVTNFKP 120 LAAETVDRDA VESAFLNSRS KAEIVDMLRK HTYAEELEQS RRIVGLDNGS GEVGGRAGCG 180 HPVGPKELLF EKVVTPSDVG KLNRLVIPKQ HAEKHFPLNT SPSASDAASK GVLLNFEDGG 240 GKVWRFRYSY WNSSQSYVLT KGWSRFVKEK GLLAGDAVSF ERSNGADRQL YIDMKHKNGE 300 LSDSQLLAHS TIRLFGVNII KDPICNIIMK HMNNPDEDNK RMRSGNEEKD LFGFQCMKKQ 360 RASPRDIVAY L* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 2e-54 | 187 | 299 | 11 | 119 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional repressor of flowering time on long day plants. Acts directly on FT expression by binding 5'-CAACA-3' and 5'-CACCTG-3 sequences. Functionally redundant with TEM2. {ECO:0000269|PubMed:18718758}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00024 | PBM | Transfer from AT1G25560 | Download |
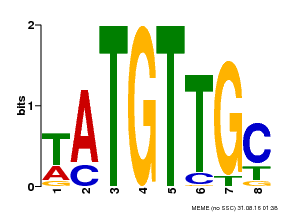
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Expressed with a circadian rhythm showing a peak at dawn. {ECO:0000269|PubMed:18718758}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002281709.2 | 1e-150 | PREDICTED: AP2/ERF and B3 domain-containing transcription factor RAV1 | ||||
| Refseq | XP_022881998.1 | 1e-151 | AP2/ERF and B3 domain-containing transcription factor RAV1 | ||||
| Swissprot | Q9C6M5 | 1e-134 | RAVL1_ARATH; AP2/ERF and B3 domain-containing transcription repressor TEM1 | ||||
| TrEMBL | A5C258 | 1e-149 | A5C258_VITVI; Uncharacterized protein | ||||
| TrEMBL | F6HFQ2 | 1e-149 | F6HFQ2_VITVI; Uncharacterized protein | ||||
| STRING | VIT_01s0011g03070.t01 | 1e-150 | (Vitis vinifera) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G13260.1 | 1e-133 | related to ABI3/VP1 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kalax.0039s0143.1.p |



