 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kalax.0022s0149.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 317aa MW: 34201.5 Da PI: 10.6695 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 31.5 | 3.5e-10 | 284 | 303 | 3 | 22 |
-SS-EEEEEEE--TT-SS-E CS
WRKY 3 DgynWrKYGqKevkgsefpr 22
D+++WrKYGqK++kgs++pr
Kalax.0022s0149.2.p 284 DEFSWRKYGQKPIKGSPHPR 303
99*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF10533 | 8.5E-16 | 236 | 280 | IPR018872 | Zn-cluster domain |
| Gene3D | G3DSA:2.20.25.80 | 1.5E-9 | 271 | 303 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 11.612 | 277 | 303 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.83E-7 | 279 | 303 | IPR003657 | WRKY domain |
| SMART | SM00774 | 0.0017 | 282 | 315 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 3.7E-6 | 284 | 303 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 317 aa Download sequence Send to blast |
MAVELMMGYR AAATENNLPA KMEEAASAGL QSVEQLIRLL SQRQQQMDRS NSNSNSNSNG 60 SCVNELSSKV ESSDLDYGAV ADVAVNKFKK MISLLDRSRT GHARFRRAPV AQKAADPVAQ 120 TSLRSEEKAS AQSSLNGGPR AIPTSTSSQR LPPLPQHHHH HHSHPVHHKI VHSANTVLTK 180 TAPKDSTATI SFSGASSFVS GLSGETESVQ PSMSSGFQFT LMSQSSGKPP LSSSLKRKCS 240 SVDDSSRCGG SSARCHCSKK RKSRVKRVVR IPAISLKMAD IPPDEFSWRK YGQKPIKGSP 300 HPRRPGVDKD LTFADF* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00450 | DAP | Transfer from AT4G24240 | Download |
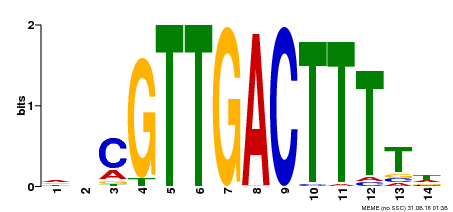
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008452580.1 | 1e-92 | PREDICTED: probable WRKY transcription factor 7 isoform X2 | ||||
| TrEMBL | A0A200QZM3 | 7e-92 | A0A200QZM3_9MAGN; DNA-binding WRKY | ||||
| STRING | XP_008452579.1 | 5e-90 | (Cucumis melo) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24240.1 | 5e-21 | WRKY DNA-binding protein 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kalax.0022s0149.2.p |



