 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kalax.0015s0146.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 385aa MW: 43314.9 Da PI: 6.5136 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 28.4 | 2.7e-09 | 311 | 346 | 20 | 55 |
HHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 20 eknrypsaeereeLAkklgLterqVkvWFqNrRake 55
+k +yps++e+ LA+++gL+++q+ +WF N+R ++
Kalax.0015s0146.1.p 311 YKWPYPSETEKVALAESTGLDQKQINNWFINQRKRH 346
4679*****************************885 PP
| |||||||
| 2 | ELK | 35.4 | 2.2e-12 | 266 | 287 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
+LK +L++KYsgyL+sLkqE+s
Kalax.0015s0146.1.p 266 DLKIHLMKKYSGYLSSLKQELS 287
69******************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 8.3E-19 | 118 | 162 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 8.1E-21 | 119 | 161 | IPR005540 | KNOX1 |
| SMART | SM01256 | 2.4E-26 | 171 | 222 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 1.4E-23 | 176 | 220 | IPR005541 | KNOX2 |
| SMART | SM01188 | 4.7E-6 | 266 | 287 | IPR005539 | ELK domain |
| PROSITE profile | PS51213 | 10.72 | 266 | 286 | IPR005539 | ELK domain |
| Pfam | PF03789 | 3.9E-9 | 266 | 287 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 13.167 | 286 | 349 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.2E-19 | 287 | 361 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 2.6E-12 | 288 | 353 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.8E-27 | 291 | 350 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 4.89E-12 | 298 | 350 | No hit | No description |
| Pfam | PF05920 | 1.5E-16 | 306 | 345 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 324 | 347 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 385 aa Download sequence Send to blast |
MEEYNNQLRE NNSSPRANFL YANPVSAASS YGIHSSTGSR STGANAASVA GQSGGSSSAF 60 QQHLYFHPDQ HNSGVKSELA ATTSQFLGQK FHQYPLGIIR ENCGNERGDH SQENDEAAAI 120 KAKIMSHPQC SNLLQAYMDC QKVGAPPQVV AQLVAAREEF ENRQRSSSSS SGKDASRDPE 180 LDQFMEAYYD MLVKYREELT RPLQEAMDFM RRIESQLNML GAGPIKSYNP SDEKSSEGVG 240 SSEEEGDNGG ETAQLPEIDP HAEDRDLKIH LMKKYSGYLS SLKQELSKKR KKGKLPKEAR 300 QKLLSWWELH YKWPYPSETE KVALAESTGL DQKQINNWFI NQRKRHWKPS EDIQFMVMDG 360 MHHQNYMDGH YQLGPDHSSY HMGP* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 281 | 290 | LKQELSKKRK |
| 2 | 287 | 291 | KKRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably binds to the DNA sequence 5'-TGAC-3'. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00675 | ChIP-seq | Transfer from GRMZM2G017087 | Download |
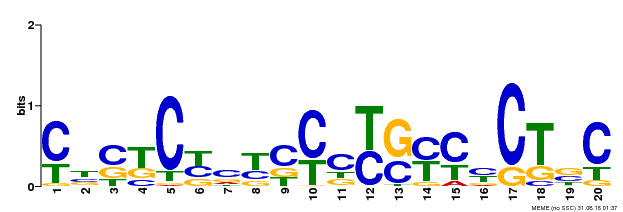
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_023903746.1 | 1e-159 | homeobox protein knotted-1-like 2 | ||||
| Swissprot | O04135 | 1e-149 | KNAP2_MALDO; Homeobox protein knotted-1-like 2 | ||||
| TrEMBL | B1P1S1 | 0.0 | B1P1S1_9MAGN; Knotted-like homeobox protein | ||||
| STRING | EOY15624 | 1e-157 | (Theobroma cacao) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 1e-100 | KNOTTED-like from Arabidopsis thaliana | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kalax.0015s0146.1.p |



