 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kaladp0577s0020.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 183aa MW: 20340.9 Da PI: 9.5861 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 46.9 | 6.5e-15 | 24 | 68 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT eE++++++a +++ + Wk+I +g ++t q++s+ qky
Kaladp0577s0020.2.p 24 RENWTDEEHDKFLEALQLFDRD-WKKIEDFVG-TKTVIQIRSHAQKY 68
789*****************77.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.96E-18 | 18 | 74 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 16.23 | 19 | 73 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-7 | 22 | 74 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.1E-18 | 22 | 71 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.1E-11 | 23 | 71 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.1E-12 | 24 | 68 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.81E-10 | 26 | 69 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0032922 | Biological Process | circadian regulation of gene expression | ||||
| GO:0043966 | Biological Process | histone H3 acetylation | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 183 aa Download sequence Send to blast |
MASSTPTDAL EKKVRKPYTI TKSRENWTDE EHDKFLEALQ LFDRDWKKIE DFVGTKTVIQ 60 IRSHAQKYFL KVQKNGTLAH VPPPRPKRKS AHPYPQKASK NADVGPKGVT TMIGSSTAGG 120 SDSSRSFPSS EVPDQDKQAP VVHGIPDFAE VYNFIGSVFD PDTKGHVQKL KEMDPINFET 180 VS* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator of evening element (EE)-containing clock-controlled genes. Forms a negative feedback loop with APRR5. Regulates the pattern of histone H3 acetylation of the TOC1 promoter. {ECO:0000269|PubMed:21205033, ECO:0000269|PubMed:21474993, ECO:0000269|PubMed:21483796, ECO:0000269|PubMed:23638299}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
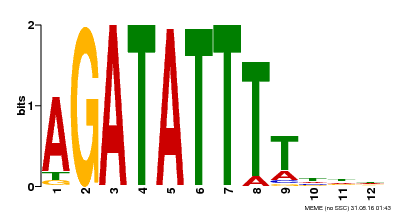
| Motif ID | Method | Source | Motif file |
| MP00338 | DAP | Transfer from AT3G09600 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Circadian-regulation. Peak of expression in the afternoon. Down-regulated by cold. {ECO:0000269|PubMed:21205033, ECO:0000269|PubMed:22902701, ECO:0000269|PubMed:23638299}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028775219.1 | 2e-98 | protein REVEILLE 8 | ||||
| Swissprot | Q8RWU3 | 4e-93 | RVE8_ARATH; Protein REVEILLE 8 | ||||
| TrEMBL | A0A371GDE8 | 1e-95 | A0A371GDE8_MUCPR; Protein REVEILLE 8 (Fragment) | ||||
| STRING | XP_007144521.1 | 5e-95 | (Phaseolus vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G09600.1 | 2e-89 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kaladp0577s0020.2.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



