 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kaladp0047s0272.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 311aa MW: 34827.7 Da PI: 6.5571 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 56.8 | 3.8e-18 | 84 | 137 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe+ +++ e++ +LA+ lgL+ rq+ +WFqNrRa++k
Kaladp0047s0272.1.p 84 KKRRLNIEQVKTLEKTFEQGNKLEPERKMQLARALGLQPRQIAIWFQNRRARWK 137
4557899**********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 129.5 | 1.4e-41 | 83 | 173 | 1 | 91 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLree 90
ekkrrl+ eqvk+LE++Fe+ +kLeperK++lar+Lglqprq+a+WFqnrRAR+ktkqlEkdy+ Lkr++da+k+ene+L +++++L++e
Kaladp0047s0272.1.p 83 EKKRRLNIEQVKTLEKTFEQGNKLEPERKMQLARALGLQPRQIAIWFQNRRARWKTKQLEKDYNILKRHFDAVKAENESLIAQNQKLHSE 172
69**************************************************************************************98 PP
HD-ZIP_I/II 91 l 91
+
Kaladp0047s0272.1.p 173 I 173
7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.25E-19 | 76 | 141 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.313 | 79 | 139 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 5.6E-18 | 82 | 143 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 1.8E-15 | 84 | 137 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.23E-17 | 84 | 140 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-19 | 86 | 146 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 1.3E-5 | 110 | 119 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 114 | 137 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.3E-5 | 119 | 135 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 8.7E-15 | 139 | 178 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0048826 | Biological Process | cotyledon morphogenesis | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 311 aa Download sequence Send to blast |
MGSNGMPFFS SNFLLQTEDD DHHHQPPTSL NPLLPSCSPQ DFHGLASLLG KRGMSFSGID 60 HICEEVMNGE EDLSDDGLSH GGEKKRRLNI EQVKTLEKTF EQGNKLEPER KMQLARALGL 120 QPRQIAIWFQ NRRARWKTKQ LEKDYNILKR HFDAVKAENE SLIAQNQKLH SEILSLRSNN 180 RSDHPESINL NKETIIDQGS CSTRSDNSSD IHLDISRTPP AIDSPPHIRP ISIPLFPTSS 240 NHHASPAAAT TTAAAQLFFS PPITGLRTDS HNTGNNSHHV KEESSFTNMF CGMEDQSGGL 300 WPWLEQQHFN * |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 131 | 139 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may act in the sucrose-signaling pathway. {ECO:0000269|PubMed:11292072}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00225 | DAP | Transfer from AT1G69780 | Download |
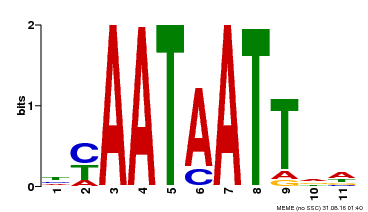
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021618165.1 | 1e-134 | homeobox-leucine zipper protein ATHB-13-like | ||||
| Refseq | XP_021618174.1 | 1e-134 | homeobox-leucine zipper protein ATHB-13-like | ||||
| Swissprot | Q8LC03 | 1e-106 | ATB13_ARATH; Homeobox-leucine zipper protein ATHB-13 | ||||
| TrEMBL | A0A2C9WNB8 | 1e-133 | A0A2C9WNB8_MANES; Uncharacterized protein | ||||
| STRING | cassava4.1_013285m | 1e-131 | (Manihot esculenta) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69780.1 | 1e-107 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kaladp0047s0272.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



