 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kaladp0042s0044.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 350aa MW: 38829.9 Da PI: 7.535 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100.1 | 1.5e-31 | 148 | 201 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
pr+rWt++LH++F++av+ LGG+e+AtPk++lelm+vk+Ltl+hvkSHLQ+YR+
Kaladp0042s0044.1.p 148 PRMRWTTTLHAHFIHAVQLLGGHERATPKSVLELMNVKDLTLAHVKSHLQMYRT 201
9****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.23E-15 | 145 | 202 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-27 | 146 | 202 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.7E-23 | 148 | 201 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.7E-7 | 149 | 200 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005618 | Cellular Component | cell wall | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 350 aa Download sequence Send to blast |
MATSASSTPP LFIPDLSLQI SQPLFSNIMS SKNDSLGIMS SGAGVATEES GSSTGSGLSH 60 ETMNGGGFQQ AEHQPTLSLG LFDHHNLLHQ QQLLLLHHNN AIFAAKSEAA ARYHQNTHHH 120 HYSPQIYGRE YKVRSSSSRT MKRSIRAPRM RWTTTLHAHF IHAVQLLGGH ERATPKSVLE 180 LMNVKDLTLA HVKSHLQMYR TVKSTDKRSG QGRNDSDLDL NANNQRAEDI ADQVNGELPS 240 PQQNAQHWGP WGSAIETSDQ QTLSTHKEAS THQPNMSHRN HRKVYDDVGV GHLVNDERKE 300 QGLECSFLST YSNNSTTTLP PQQHQGLLNL EFTLGRPSWE LERQHAAGY* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 1e-15 | 149 | 203 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 1e-15 | 149 | 203 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 1e-15 | 149 | 203 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 1e-15 | 149 | 203 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 1e-15 | 149 | 203 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 1e-15 | 149 | 203 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 1e-15 | 149 | 203 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 1e-15 | 149 | 203 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00107 | PBM | Transfer from AT5G42630 | Download |
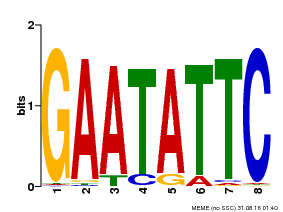
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42630.1 | 1e-54 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kaladp0042s0044.1.p |



