 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kaladp0003s0022.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 1001aa MW: 109445 Da PI: 6.1378 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 90.9 | 9.9e-29 | 603 | 653 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isl++l++yF++++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rki+++
Kaladp0003s0022.1.p 603 EKTISLDVLQQYFAGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKINKV 653
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.598 | 592 | 673 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 6.7E-26 | 605 | 653 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 9.48E-21 | 902 | 990 | No hit | No description |
| SMART | SM00666 | 5.0E-24 | 905 | 987 | IPR000270 | PB1 domain |
| PROSITE profile | PS51745 | 23.364 | 905 | 987 | IPR000270 | PB1 domain |
| CDD | cd06407 | 4.34E-35 | 906 | 986 | No hit | No description |
| Pfam | PF00564 | 7.1E-19 | 906 | 986 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 1.7E-23 | 906 | 986 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1001 aa Download sequence Send to blast |
MAEPDDDKAS SRFPLHSHHQ SRSSRDPPPD SHHLMDLDLD LDSFWPLDQI SFATNPMSPL 60 LISSSDQPFS PLWAFSDADD DNKPSSASAK QAALSAAAAF RLPDYPRFLA LKSAYTDTTN 120 GIPADNDEKR RMQSLPPSSP LWGLEPDVSC VIKEKMTQAL RYFKESTEQH VLAQVWAPVK 180 NGGRYVLTTS EQPFVLDPHS SGLNRYRMVS LMYMFCVDEE SDGELGLPGR VFRQKLPEWT 240 PNVQYYSSQE YSRLDHALHY NVQGTLALPV FESSGQTCVG VVELILTSQK INYAPEVDKV 300 CKALEAVNLK SSNILDHPKP QIRNEGRQKA LAEILEVLSV VCESLKLPLA QTWVPCRHRN 360 VLADGGGLRK SCSSFDGSCM GQVCMSTSDV TFYVVDAHLW GFRDACAEHH LQRGQGVAGR 420 AFLSQSSCFC GDITKFSKTE YPLVHYARLF GVTSCFAICL CSSLTGNDEY ILEFFLPPTV 480 IDARDQHTLL NSLLETMKQH FRSLRVASGR NLGEDMIEIV EGSADEKLLE SVRRSHSASP 540 PAPDTTNGRE TCDKISTVNL VLEEANGSNN AMNAAGLGGI TSGGTLMGNT VTKKPERKRG 600 KTEKTISLDV LQQYFAGSLK DAAKSLGVCP TTMKRICRQH GISRWPSRKI NKVNRSLTKL 660 KRVIESVQGS ESTFGLTSLS PTPLPAAGCS MSWAANANGS NLPNSPGLRT AGLQGMKSES 720 PECKSPISDG KDGNGVHLPG SKTLDPEGCP NEENEYIPKI SKTSSGSREG SAGSPTSHGS 780 CQGSAPHETN VAQYSKTPEL CEPYINIEAP LRNHQGKELD LAMTNFVPDT RLSTQAQEPF 840 EGLLVADVGS SKNLRNLCPS IADAHIEDRH PECSWTNSHV PIPASEPATT ANPTPHLTGR 900 RDSVTLTIKA TYKDDIIRFR FLSNSGIVEL REEVAKRLKL EVGTFDIKYL DDDHELVLIV 960 CDADLQECFD VSRSSGSLMI RLSVHDITTN LGSSCESTGE * |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
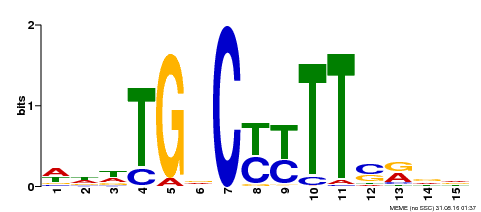
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010659716.1 | 0.0 | PREDICTED: protein NLP6 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | F6HU52 | 0.0 | F6HU52_VITVI; Uncharacterized protein | ||||
| STRING | VIT_02s0025g02070.t01 | 0.0 | (Vitis vinifera) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kaladp0003s0022.1.p |



