 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | kfl00198_0060 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Klebsormidiophyceae; Klebsormidiales; Klebsormidiaceae; Klebsormidium
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 357aa MW: 38127.8 Da PI: 8.8022 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 14.2 | 0.00013 | 60 | 82 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+ C+ C Fs + +L++H+ H
kfl00198_0060 60 HICEVCAIAFSKPAHLRQHLVIH 82
68*****************9887 PP
| |||||||
| 2 | zf-C2H2 | 21 | 9e-07 | 88 | 112 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++Cp C ++F+r+++L+rH+ +H
kfl00198_0060 88 FPCPhaGCARTFKRQDHLRRHLLSH 112
89********************999 PP
| |||||||
| 3 | zf-C2H2 | 12.7 | 0.00038 | 118 | 141 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirtH 23
+Cp C+ ++s ++nL+ H ++H
kfl00198_0060 118 PCPipGCPVTCSLPDNLRAHVKRH 141
69999******************9 PP
| |||||||
| 4 | zf-C2H2 | 17 | 1.7e-05 | 175 | 200 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
+ C+ C+k+F+ +s+L+ H+ + H
kfl00198_0060 175 HMCQepGCSKRFKFRSLLTAHQASvH 200
578888****************9988 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.160.60 | 5.0E-10 | 57 | 81 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.68E-11 | 58 | 112 | No hit | No description |
| SMART | SM00355 | 0.065 | 60 | 82 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.422 | 60 | 87 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 62 | 82 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.6E-14 | 82 | 114 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| Pfam | PF13894 | 0.0013 | 88 | 112 | No hit | No description |
| SMART | SM00355 | 4.0E-4 | 88 | 112 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.471 | 88 | 113 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 90 | 112 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.024 | 117 | 141 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 119 | 141 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 3.8 | 146 | 171 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.141 | 175 | 200 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.093 | 175 | 200 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-4 | 175 | 196 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 177 | 200 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.081 | 262 | 287 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 3.8E-4 | 262 | 285 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 264 | 287 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 1.6 | 293 | 318 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 295 | 318 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 357 aa Download sequence Send to blast |
MGPVKHEDGG DSTDGGDGTD SGRLRKKSLR GASGKAACSL RLEQQQPEEE QVGTSSERQH 60 ICEVCAIAFS KPAHLRQHLV IHTGQRPFPC PHAGCARTFK RQDHLRRHLL SHSQPVLPCP 120 IPGCPVTCSL PDNLRAHVKR HQAACFRCSE PECTFETAFK RQLWEHTRSC HASGHMCQEP 180 GCSKRFKFRS LLTAHQASVH GFTANRAAPL VSNIGASSNT NRGPGCKDAQ ECSTPRPGDS 240 GRSSSGAATS TDEEAGSGPP GLACPVPGCT AQFRLRRSLK QHLTVAHLKE ARHPCSTAGC 300 PASFFTEADR ARHSATKHPP AGVGVATGGA SQSGMQAARQ RPSIASILDQ PTFADPI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tf6_A | 8e-12 | 58 | 202 | 11 | 162 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| 1tf6_D | 8e-12 | 58 | 202 | 11 | 162 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
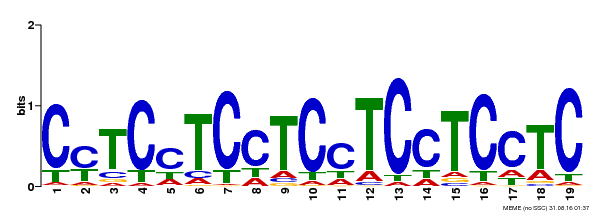
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A1Y1I138 | 0.0 | A0A1Y1I138_KLENI; Uncharacterized protein | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP2981 | 16 | 27 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 2e-18 | transcription factor IIIA | ||||



